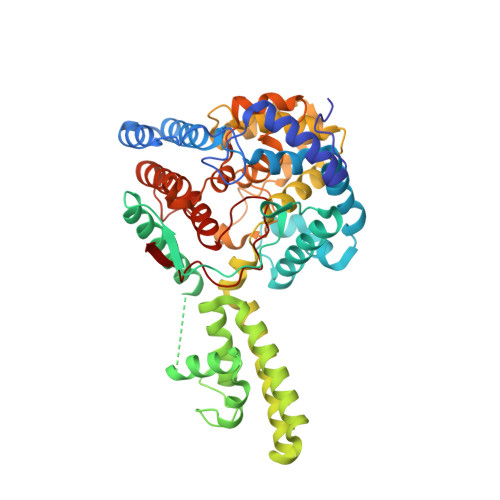
The Crystal Structure of the Hazara Virus Nucleocapsid Protein.
Surtees, R., Ariza, A., Punch, E.K., Trinh, C.H., Dowall, S.D., Hewson, R., Hiscox, J.A., Barr, J.N., Edwards, T.A.(2015) BMC Struct Biol 15: 24
- PubMed: 26715309 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/s12900-015-0051-3
- Primary Citation Related Structures:
5A97 - PubMed Abstract:
Hazara virus (HAZV) is a member of the Bunyaviridae family of segmented negative stranded RNA viruses, and shares the same serogroup as Crimean-Congo haemorrhagic fever virus (CCHFV). CCHFV is responsible for fatal human disease with a mortality rate approaching 30 %, which has an increased recent incidence within southern Europe. There are no preventative or therapeutic treatments for CCHFV-mediated disease, and thus CCHFV is classified as a hazard group 4 pathogen. In contrast HAZV is not associated with serious human disease, although infection of interferon receptor knockout mice with either CCHFV or HAZV results in similar disease progression. To characterise further similarities between HAZV and CCHFV, and support the use of HAZV as a model for CCHFV infection, we investigated the structure of the HAZV nucleocapsid protein (N) and compared it to CCHFV N. N performs an essential role in the viral life cycle by encapsidating the viral RNA genome, and thus, N represents a potential therapeutic target. We present the purification, crystallisation and crystal structure of HAZV N at 2.7 Å resolution. HAZV N was expressed as an N-terminal glutathione S-transferase (GST) fusion protein then purified using glutathione affinity chromatography followed by ion-exchange chromatography. HAZV N crystallised in the P212121 space group with unit cell parameters a = 64.99, b = 76.10, and c = 449.28 Å. HAZV N consists of a globular domain formed mostly of alpha helices derived from both the N- and C-termini, and an arm domain comprising two long alpha helices. HAZV N has a similar overall structure to CCHFV N, with their globular domains superposing with an RMSD = 0.70 Å, over 368 alpha carbons that share 59 % sequence identity. Four HAZV N monomers crystallised in the asymmetric unit, and their head-to-tail assembly reveals a potential interaction site between monomers. The crystal structure of HAZV N reveals a close similarity to CCHFV N, supporting the use of HAZV as a model for CCHFV. Structural similarity between the N proteins should facilitate study of the CCHFV and HAZV replication cycles without the necessity of working under containment level 4 (CL-4) conditions.
- Public Health England, Porton Down, Salisbury, Wiltshire, SP4 0JG, UK. bs06ras@leeds.ac.uk.
Organizational Affiliation: