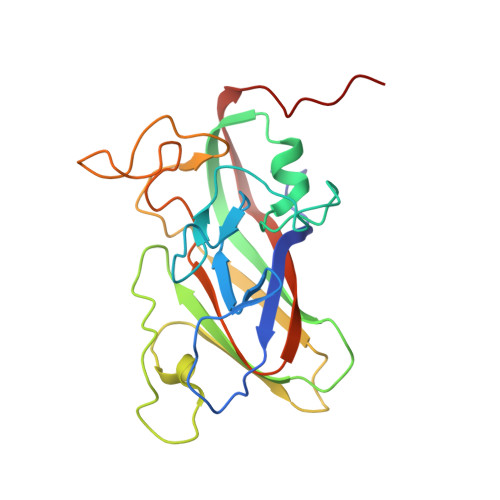
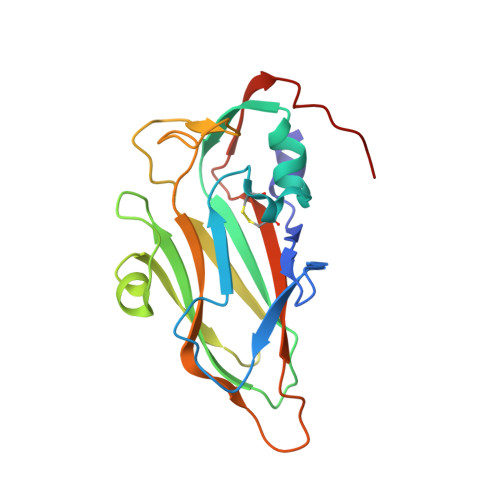
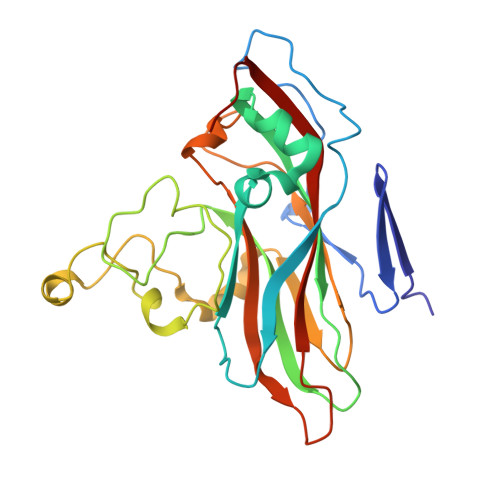
Structure and Genome Release Mechanism of Human Cardiovirus Saffold Virus-3.
Mullapudi, E., Novacek, J., Palkova, L., Kulich, P., Lindberg, A.M., Van Kuppeveld, F.J.M., Plevka, P.(2016) J Virol 90: 7628
- PubMed: 27279624 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.00746-16
- Primary Citation Related Structures:
5A8F, 5CFC, 5CFD - PubMed Abstract:
In order to initiate an infection, viruses need to deliver their genomes into cells. This involves uncoating the genome and transporting it to the cytoplasm. The process of genome delivery is not well understood for nonenveloped viruses. We address this gap in our current knowledge by studying the uncoating of the nonenveloped human cardiovirus Saffold virus 3 (SAFV-3) of the family Picornaviridae SAFVs cause diseases ranging from gastrointestinal disorders to meningitis. We present a structure of a native SAFV-3 virion determined to 2.5 Å by X-ray crystallography and an 11-Å-resolution cryo-electron microscopy reconstruction of an "altered" particle that is primed for genome release. The altered particles are expanded relative to the native virus and contain pores in the capsid that might serve as channels for the release of VP4 subunits, N termini of VP1, and the RNA genome. Unlike in the related enteroviruses, pores in SAFV-3 are located roughly between the icosahedral 3- and 5-fold axes at an interface formed by two VP1 and one VP3 subunit. Furthermore, in native conditions many cardioviruses contain a disulfide bond formed by cysteines that are separated by just one residue. The disulfide bond is located in a surface loop of VP3. We determined the structure of the SAFV-3 virion in which the disulfide bonds are reduced. Disruption of the bond had minimal effect on the structure of the loop, but it increased the stability and decreased the infectivity of the virus. Therefore, compounds specifically disrupting or binding to the disulfide bond might limit SAFV infection. A capsid assembled from viral proteins protects the virus genome during transmission from one cell to another. However, when a virus enters a cell the virus genome has to be released from the capsid in order to initiate infection. This process is not well understood for nonenveloped viruses. We address this gap in our current knowledge by studying the genome release of Human Saffold virus 3 Saffold viruses cause diseases ranging from gastrointestinal disorders to meningitis. We show that before the genome is released, the Saffold virus 3 particle expands, and holes form in the previously compact capsid. These holes serve as channels for the release of the genome and small capsid proteins VP4 that in related enteroviruses facilitate subsequent transport of the virus genome into the cell cytoplasm.
- Central European Institute of Technology, Masaryk University, Brno, Czech Republic.
Organizational Affiliation: