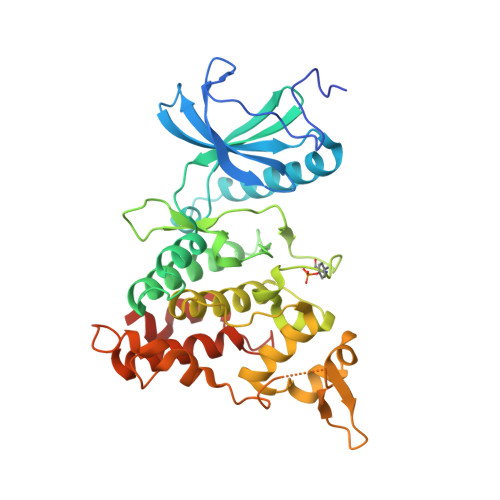
Probing the ATP-Binding Pocket of Protein Kinase Dyrk1A with Benzothiazole Fragment Molecules
Rothweiler, U., Stensen, W., Brandsdal, B.O., Isaksson, J., Leeson, F.A., Eugh, R.A., Mjoen Svendsen, J.S.(2016) J Med Chem 59: 9814
- PubMed: 27736065 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.6b01086
- Primary Citation Related Structures:
5A3X, 5A4E, 5A4L, 5A4Q, 5A4T, 5A54 - PubMed Abstract:
DYRK1A has emerged as a potential target for therapies of Alzheimer's disease using small molecules. On the basis of the observation of selective DYRK1A inhibition by firefly d-luciferin, we have explored static and dynamic structural properties of fragment sized variants of the benzothiazole scaffold with respect to DYRK1A using X-ray crystallography and NMR techniques. The compounds have excellent ligand efficiencies and show a remarkable diversity of binding modes in dynamic equilibrium. Binding geometries are determined in part by interactions often considered "weak", including "orthogonal multipolar" types represented by, for example, F-CO, sulfur-aromatic, and halogen-aromatic interactions, together with hydrogen bonds that are modulated by variation of electron withdrawing groups. These studies show how the benzothiazole scaffold is highly promising for the development of therapeutic DYRK1A inhibitors. In addition, the subtleties of the binding interactions, including dynamics, show how full structural studies are required to fully interpret the essential physical determinants of binding.
- The Norwegian Structural Biology Centre, Department of Chemistry, UiT The Arctic University of Norway , N-9037 Tromsø, Norway.
Organizational Affiliation: