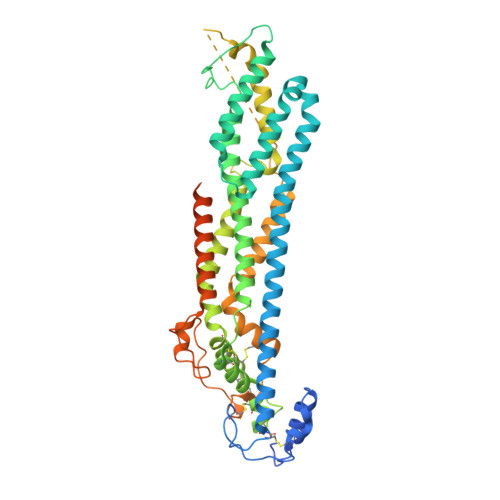
Structural Aspects of N-Glycosylations and the C-terminal Region in Human Glypican-1.
Awad, W., Adamczyk, B., Ornros, J., Karlsson, N.G., Mani, K., Logan, D.T.(2015) J Biological Chem 290: 22991-23008
- PubMed: 26203194 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.660878
- Primary Citation Related Structures:
4YWT - PubMed Abstract:
Glypicans are multifunctional cell surface proteoglycans involved in several important cellular signaling pathways. Glypican-1 (Gpc1) is the predominant heparan sulfate proteoglycan in the developing and adult human brain. The two N-linked glycans and the C-terminal domain that attach the core protein to the cell membrane are not resolved in the Gpc1 crystal structure. Therefore, we have studied Gpc1 using crystallography, small angle x-ray scattering, and chromatographic approaches to elucidate the composition, structure, and function of the N-glycans and the C terminus and also the topology of Gpc1 with respect to the membrane. The C terminus is shown to be highly flexible in solution, but it orients the core protein transverse to the membrane, directing a surface evolutionarily conserved in Gpc1 orthologs toward the membrane, where it may interact with signaling molecules and/or membrane receptors on the cell surface, or even the enzymes involved in heparan sulfate substitution in the Golgi apparatus. Furthermore, the N-glycans are shown to extend the protein stability and lifetime by protection against proteolysis and aggregation.
- From the Department of Biochemistry and Structural Biology, Centre for Molecular Protein Science, Lund University, Box 124, SE-221 00 Lund.
Organizational Affiliation: