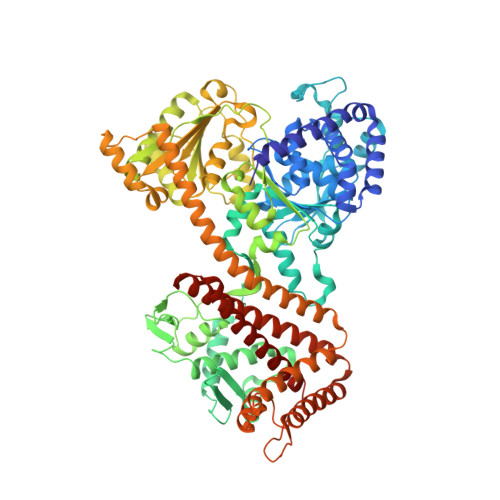
Conformational Changes of the Clamp of the Protein Translocation ATPase SecA.
Chen, Y., Bauer, B.W., Rapoport, T.A., Gumbart, J.C.(2015) J Mol Biology 427: 2348-2359
- PubMed: 25982945 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2015.05.003
- Primary Citation Related Structures:
4YS0 - PubMed Abstract:
Post-translational protein translocation across the bacterial plasma membrane is mediated by the interplay of the SecA ATPase and the protein-conducting SecY channel. SecA consists of several domains, including two nucleotide-binding domains (NBD1 and NBD2), a polypeptide cross-linking domain (PPXD), a helical scaffold domain (HSD), and a helical wing domain (HWD). PPXD, HSD, and NBD2 form a clamp that positions the polypeptide substrate above the channel so that it can be pushed into the channel by a two-helix finger of the HSD. How the substrate is accommodated in the clamp during translocation is unclear. Here, we report a crystal structure of Thermotoga maritima SecA at 1.9 Å resolution. Structural analysis and free-energy calculations indicate that the new structure represents an intermediate state during the transition of the clamp from an open to a closed conformation. Molecular dynamics simulations show that closure of the clamp occurs in two phases, an initial movement of PPXD, HSD, and HWD as a unit, followed by a movement of PPXD alone toward NBD2. Simulations in the presence of a polypeptide chain show that the substrate associates with the back of the clamp by dynamic hydrogen bonding and that the clamp is laterally closed by a conserved loop of the PPXD. Mutational disruption of clamp opening or closure abolishes protein translocation. These results suggest how conformational changes of SecA allow substrate binding and movement during protein translocation.
- Howard Hughes Medical Institute and Department of Cell Biology, Harvard Medical School, 240 Longwood Avenue, Boston, MA 02115, USA.
Organizational Affiliation: