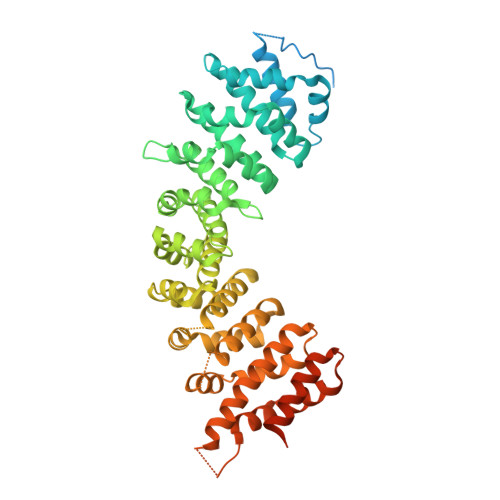
Probing formation of cargo/importin-alpha transport complexes in plant cells using a pathogen effector.
Wirthmueller, L., Roth, C., Fabro, G., Caillaud, M.C., Rallapalli, G., Asai, S., Sklenar, J., Jones, A.M., Wiermer, M., Jones, J.D., Banfield, M.J.(2015) Plant J 81: 40-52
- PubMed: 25284001 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/tpj.12691
- Primary Citation Related Structures:
4TNM - PubMed Abstract:
Importin-αs are essential adapter proteins that recruit cytoplasmic proteins destined for active nuclear import to the nuclear transport machinery. Cargo proteins interact with the importin-α armadillo repeat domain via nuclear localization sequences (NLSs), short amino acids motifs enriched in Lys and Arg residues. Plant genomes typically encode several importin-α paralogs that can have both specific and partially redundant functions. Although some cargos are preferentially imported by a distinct importin-α it remains unknown how this specificity is generated and to what extent cargos compete for binding to nuclear transport receptors. Here we report that the effector protein HaRxL106 from the oomycete pathogen Hyaloperonospora arabidopsidis co-opts the host cell's nuclear import machinery. We use HaRxL106 as a probe to determine redundant and specific functions of importin-α paralogs from Arabidopsis thaliana. A crystal structure of the importin-α3/MOS6 armadillo repeat domain suggests that five of the six Arabidopsis importin-αs expressed in rosette leaves have an almost identical NLS-binding site. Comparison of the importin-α binding affinities of HaRxL106 and other cargos in vitro and in plant cells suggests that relatively small affinity differences in vitro affect the rate of transport complex formation in vivo. Our results suggest that cargo affinity for importin-α, sequence variation at the importin-α NLS-binding sites and tissue-specific expression levels of importin-αs determine formation of cargo/importin-α transport complexes in plant cells.
- The Sainsbury Laboratory, Norwich Research Park, Norwich, NR4 7UH, UK; Department of Biological Chemistry, John Innes Centre, Norwich Research Park, Norwich, NR4 7UH, UK.
Organizational Affiliation: