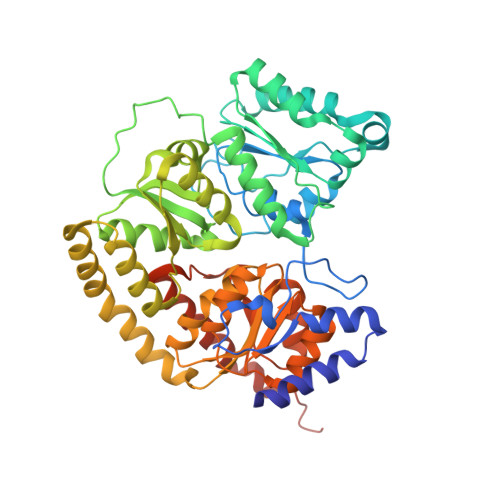
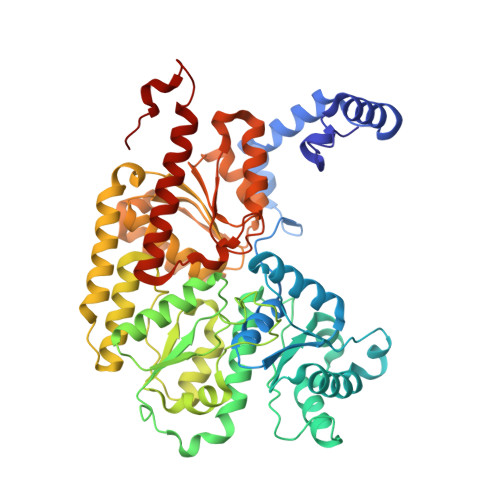
Ligand binding to the FeMo-cofactor: structures of CO-bound and reactivated nitrogenase.
Spatzal, T., Perez, K.A., Einsle, O., Howard, J.B., Rees, D.C.(2014) Science 345: 1620-1623
- PubMed: 25258081 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.1256679
- Primary Citation Related Structures:
4TKU, 4TKV - PubMed Abstract:
The mechanism of nitrogenase remains enigmatic, with a major unresolved issue concerning how inhibitors and substrates bind to the active site. We report a crystal structure of carbon monoxide (CO)-inhibited nitrogenase molybdenum-iron (MoFe)-protein at 1.50 angstrom resolution, which reveals a CO molecule bridging Fe2 and Fe6 of the FeMo-cofactor. The μ2 binding geometry is achieved by replacing a belt-sulfur atom (S2B) and highlights the generation of a reactive iron species uncovered by the displacement of sulfur. The CO inhibition is fully reversible as established by regain of enzyme activity and reappearance of S2B in the 1.43 angstrom resolution structure of the reactivated enzyme. The substantial and reversible reorganization of the FeMo-cofactor accompanying CO binding was unanticipated and provides insights into a catalytically competent state of nitrogenase.
- Howard Hughes Medical Institute and Division of Chemistry and Chemical Engineering, MailCode 114-96, California Institute of Technology, Pasadena, CA 91125, USA. spatzal@caltech.edu dcrees@caltech.edu.
Organizational Affiliation: