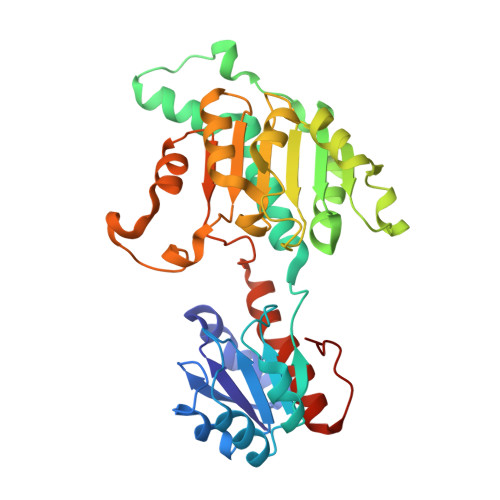
Crystal structure and thermodynamic properties of d-lactate dehydrogenase from Lactobacillus jensenii.
Kim, S., Gu, S.A., Kim, Y.H., Kim, K.J.(2014) Int J Biol Macromol 68C: 151-157
- PubMed: 24794195 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2014.04.048
- Primary Citation Related Structures:
4PRK, 4PRL - PubMed Abstract:
The thermostable d-lactate dehydrogenase from Lactobacillus jensenii (Ljd-LDH) is a key enzyme in the production of the d-form of lactic acid from pyruvate concomitant with the oxidation of NADH to NAD(+). The polymers of d-lactic acid are used as biodegradable bioplastics. The crystal structures of Ljd-LDH and in complex with NAD(+) were determined at 2.13 and 2.60Å resolutions, respectively. The Ljd-LDH monomer consists of the N-terminal substrate-binding domain and the C-terminal NAD-binding domain. The Ljd-LDH forms a homodimeric structure, and the C-terminal NAD-binding domain mostly enables the dimerization of the enzyme. The NAD cofactor is bound to the GxGxxG NAD-binding motif located between the two domains. Structural comparisons of Ljd-LDH with other d-LDHs reveal that Ljd-LDH has unique amino acid residues at the linker region, which indicates that the open-close dynamics of Ljd-LDH might be different from that of other d-LDHs. Moreover, thermostability experiments showed that the T50(10) value of Ljd-LDH (54.5°C) was much higher than the commercially available d-lactate dehydrogenase (42.7°C). In addition, Ljd-LDH has at least a 7°C higher denaturation temperature compared to commercially available d-LDHs.
- School of Life Sciences, KNU Creative BioResearch Group (BK21 plus program), Kyungpook National University, Daehak-ro 80, Buk-ku, Daegu 702-701, Republic of Korea; School of Nano-Bioscience and Chemical Engineering, Ulsan National Institute of Science and Technology (UNIST), Ulsan 689-798, Republic of Korea.
Organizational Affiliation: