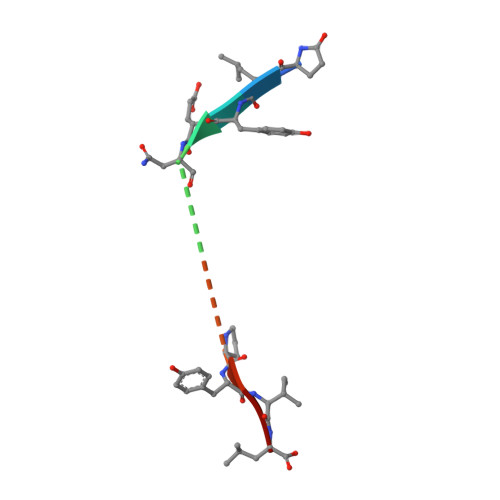
Revisiting the structure of the Vps10 domain of human sortilin and its interaction with neurotensin.
Quistgaard, E.M., Grftehauge, M.K., Madsen, P., Pallesen, L.T., Christensen, B., Srensen, E.S., Nissen, P., Petersen, C.M., Thirup, S.S.(2014) Protein Sci 23: 1291-1300
- PubMed: 24985322 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2512
- Primary Citation Related Structures:
4PO7 - PubMed Abstract:
Sortilin is a multifunctional receptor involved in sorting and apoptosis. We have previously reported a 2.0-Å structure of the Vps10 ectodomain in complex with one of its ligands, the tridecapeptide neurotensin. Here we set out to further characterize the structural properties of sortilin and its interaction with neurotensin. To this end, we have determined a new 2.7 Å structure using a crystal grown with a 10-fold increased concentration of neurotensin. Here a second peptide fragment was observed within the Vps10 β-propeller, which may in principle either represent a second molecule of neurotensin or the N-terminal part of the molecule bound at the previously identified binding site. However, in vitro binding experiments strongly favor the latter hypothesis. Neurotensin thus appears to bind with a 1:1 stoichiometry, and whereas the N-terminus does not bind on its own, it enhances the affinity in context of full-length neurotensin. We conclude that the N-terminus of neurotensin probably functions as an affinity enhancer for binding to sortilin by engaging the second binding site. Crystal packing differs partly from the previous structure, which may be due to variations in the degree and pattern of glycosylations. Consequently, a notable hydrophobic loop, not modeled previously, could now be traced. A computational analysis suggests that this and a neighboring loop may insert into the membrane and thus restrain movement of the Vps10 domain. We have, furthermore, mapped all N-linked glycosylations of CHO-expressed human sortilin by mass spectrometry and find that their locations are compatible with membrane insertion of the hydrophobic loops.
- Department of Molecular Biology and Genetics, MIND Centre, Aarhus University, Gustav Wieds Vej 10C, DK 8000, Aarhus C, Denmark; Department of Medical Biochemistry and Biophysics, Karolinska Institute, 17177, Stockholm, Sweden.
Organizational Affiliation: