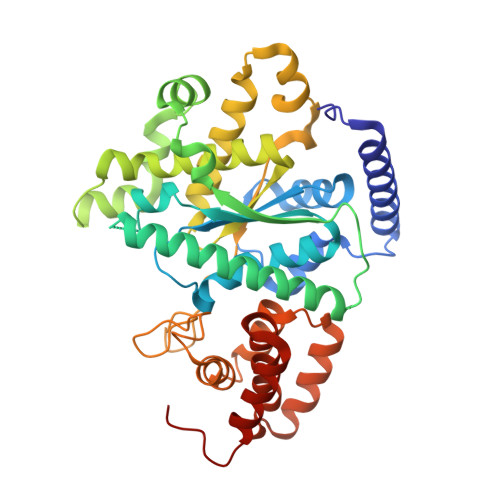
An in Vitro Peptide Complementation Assay for CYT-18-Dependent Group I Intron Splicing Reveals a New Role for the N-Terminus.
Geng, C., Paukstelis, P.J.(2014) Biochemistry 53: 1311-1319
- PubMed: 24520960 Search on PubMed
- DOI: https://doi.org/10.1021/bi401614h
- Primary Citation Related Structures:
4OJM - PubMed Abstract:
The mitochondrial tyrosyl tRNA synthetase from Neurospora crassa (CYT-18 protein) is a bifunctional group I intron splicing cofactor. CYT-18 is capable of splicing multiple group I introns from a wide variety of sources by stabilizing the catalytically active intron structures. CYT-18 and mt TyrRSs from related fungal species have evolved to assist in group I intron splicing in part by the accumulation of three N-terminal domain insertions. Biochemical and structural analysis indicate that the N-terminal insertions serve primarily to create a structure-stabilizing scaffold for critical tertiary interactions between the two major RNA domains of group I introns. Previous studies concluded that the primarily α-helical N-terminal insertion, H0, contributes to protein stability and is necessary for splicing the N. crassa ND1 intron but is dispensable for splicing the N. crassa mitochondrial LSU intron. Here, we show that CYT-18 with a complete H0 deletion retains residual ND1 intron splicing activity and that addition of the missing N-terminus in trans is capable of restoring a significant portion of its splicing activity. The development of this peptide complementation assay has allowed us to explore important characteristics of the CYT-18/group I intron interaction including the stoichiometry of H0 in intron splicing and the importance of specific H0 residues. Evaluation of truncated H0 peptides in this assay and a re-examination of the CYT-18 crystal structure suggest a previously unknown structural role of the first five N-terminal residues of CYT-18. These residues interact directly with another splicing insertion, making H0 a central structural element responsible for connecting all three N-terminal splicing insertions.
- University of Maryland , Department of Chemistry and Biochemistry, Center for Biomolecular Structure and Organization, College Park, Maryland 20742, United States.
Organizational Affiliation: