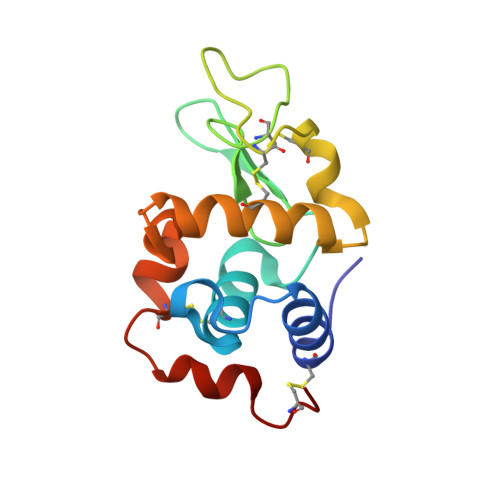
Peculiar features in the crystal structure of the adduct formed between cis-PtI2(NH3)2 and hen egg white lysozyme.
Messori, L., Marzo, T., Gabbiani, C., Valdes, A.A., Quiroga, A.G., Merlino, A.(2013) Inorg Chem 52: 13827-13829
- PubMed: 24256441 Search on PubMed
- DOI: https://doi.org/10.1021/ic402611m
- Primary Citation Related Structures:
4MR1 - PubMed Abstract:
The reactivity of cis-diamminediiodidoplatinum(II), cis-PtI2(NH3)2, the iodo analogue of cisplatin, with hen egg white lysozyme (HEWL) was investigated by electrospray ionization mass spectrometry and X-ray crystallography. Interestingly, the study compound forms a stable 1:1 protein adduct for which the crystal structure was solved at 1.99 Å resolution. In this adduct, the Pt(II) center, upon release of one ammonia ligand, selectively coordinates to the imidazole of His15. Both iodide ligands remain bound to platinum, with this being a highly peculiar and unexpected feature. Notably, two equivalent modes of Pt(II) binding are possible that differ only in the location of I atoms with respect to ND1 of His15. The structure of the adduct was compared with that of HEWL-cisplatin, previously described; differences are stressed and their important mechanistic implications discussed.
- Department of Chemistry, University of Florence , Via della Lastruccia 3, 50019 Sesto Fiorentino, Italy.
Organizational Affiliation: