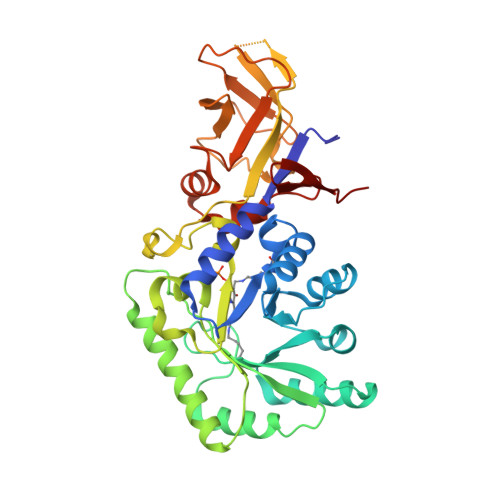
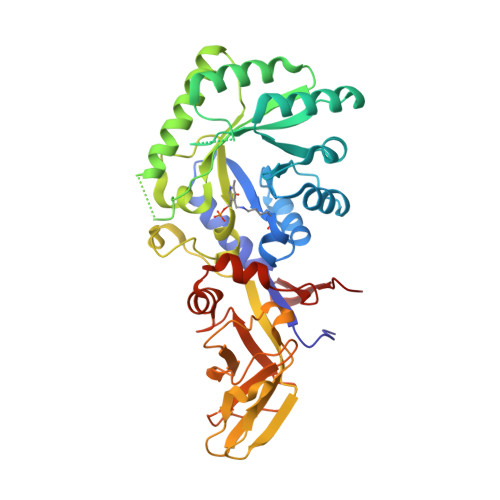
Structural and biochemical analyses of alanine racemase from the multidrug-resistant Clostridium difficile strain 630.
Asojo, O.A., Nelson, S.K., Mootien, S., Lee, Y., Rezende, W.C., Hyman, D.A., Matsumoto, M.M., Reiling, S., Kelleher, A., Ledizet, M., Koski, R.A., Anthony, K.G.(2014) Acta Crystallogr D Biol Crystallogr 70: 1922-1933
- PubMed: 25004969 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1399004714009419
- Primary Citation Related Structures:
4LUS, 4LUT, 4LUY - PubMed Abstract:
Clostridium difficile, a Gram-positive, spore-forming anaerobic bacterium, is the leading cause of infectious diarrhea among hospitalized patients. C. difficile is frequently associated with antibiotic treatment, and causes diseases ranging from antibiotic-associated diarrhea to life-threatening pseudomembranous colitis. The severity of C. difficile infections is exacerbated by the emergence of hypervirulent and multidrug-resistant strains, which are difficult to treat and are often associated with increased mortality rates. Alanine racemase (Alr) is a pyridoxal-5'-phosphate (PLP)-dependent enzyme that catalyzes the reversible racemization of L- and D-alanine. Since D-alanine is an essential component of the bacterial cell-wall peptidoglycan, and there are no known Alr homologs in humans, this enzyme is being tested as an antibiotic target. Cycloserine is an antibiotic that inhibits Alr. In this study, the catalytic properties and crystal structures of recombinant Alr from the virulent and multidrug-resistant C. difficile strain 630 are presented. Three crystal structures of C. difficile Alr (CdAlr), corresponding to the complex with PLP, the complex with cycloserine and a K271T mutant form of the enzyme with bound PLP, are presented. The structures are prototypical Alr homodimers with two active sites in which the cofactor PLP and cycloserine are localized. Kinetic analyses reveal that the K271T mutant CdAlr has the highest catalytic constants reported to date for any Alr. Additional studies are needed to identify the basis for the high catalytic activity. The structural and activity data presented are first steps towards using CdAlr for the development of structure-based therapeutics for C. difficile infections.
- National School of Tropical Medicine, Department of Pediatrics, Baylor College of Medicine, Houston, TX 77030, USA.
Organizational Affiliation: