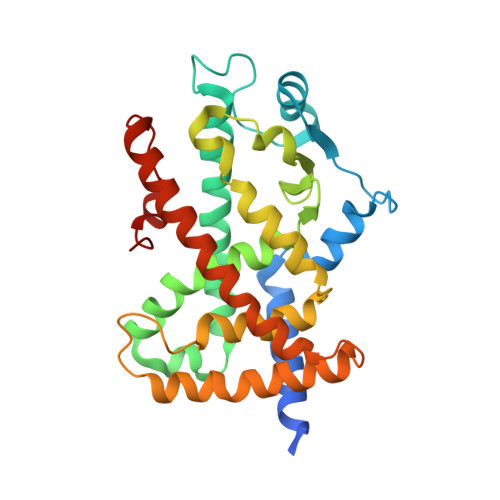
Resveratrol and Its Metabolites Bind to PPARs.
Calleri, E., Pochetti, G., Dossou, K.S., Laghezza, A., Montanari, R., Capelli, D., Prada, E., Loiodice, F., Massolini, G., Bernier, M., Moaddel, R.(2014) Chembiochem 15: 1154-1160
- PubMed: 24796862
- DOI: https://doi.org/10.1002/cbic.201300754
- Primary Citation of Related Structures:
4JAZ - PubMed Abstract:
Resveratrol, a modulator of several signaling proteins, can exert off-target effects involving the peroxisome proliferator-activated receptor (PPAR) transcription factors. However, evidence for the direct interaction between this polyphenol and PPARs is lacking. Here, we addressed the hypothesis that resveratrol and its metabolites control aspects of PPAR transcriptional activity through direct interaction with PPARs. Bioaffinity chromatographic studies with the immobilized ligand-binding domains (LBDs) of PPARγ and PPARα and isothermal titration calorimetry allowed the binding affinities of resveratrol, resveratrol 3-O-glucuronide, resveratrol 4-O-glucuronide, and resveratrol 3-O-sulfate to both PPAR-LBDs to be determined. Interaction of resveratrol, resveratrol 3-O-glucuronide, and resveratrol 4-O-glucuronide with PPARγ-LBD occurred with binding affinities of 1.4, 1.1, and 0.8 μM, respectively, although only resveratrol bound to the PPARα-LBD with a binding affinity of 2.7 μM. Subsequently, X-ray crystallographic studies were carried out to characterize resveratrol binding to the PPARγ-LBD at the molecular level. The electron density map from the crystal structure of the complex between PPARγ-LBD and resveratrol revealed the presence of one molecule of resveratrol bound to the LBD of PPARγ, with the ligand occupying a position close to that of other known PPARγ ligands. Transactivation assays were also performed in HepG2 cells, with the results showing that resveratrol was not a PPAR agonist but instead was able to displace rosiglitazone from PPARγ and Wy-14643 from PPARα with IC50 values of (27.4±1.8) μM and (31.7±2.5) μM, respectively. We propose that resveratrol acts as a PPAR antagonist through its direct interaction with PPARγ and PPARα.
- Dipartimento di Scienze del Farmaco, Università degli Studi di Pavia, 27100 Pavia, Italy.
Organizational Affiliation: