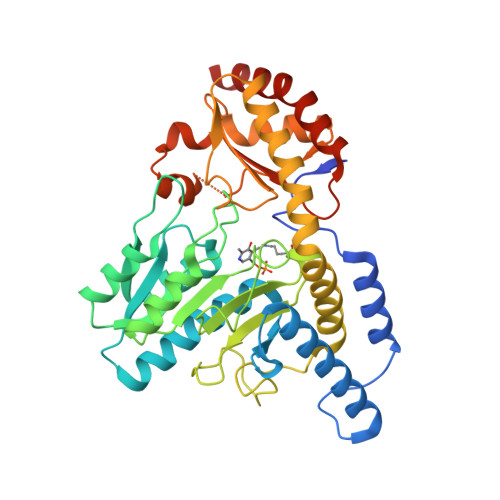
The cysteine desulfurase IscS of Mycobacterium tuberculosis is involved in iron-sulfur cluster biogenesis and oxidative stress defence.
Rybniker, J., Pojer, F., Marienhagen, J., Kolly, G.S., Chen, J.M., van Gumpel, E., Hartmann, P., Cole, S.T.(2014) Biochem J 459: 467-478
- PubMed: 24548275 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20130732
- Primary Citation Related Structures:
4ISY - PubMed Abstract:
The complex multiprotein systems for the assembly of protein-bound iron-sulfur (Fe-S) clusters are well defined in Gram-negative model organisms. However, little is known about Fe-S cluster biogenesis in other bacterial species. The ISC (iron-sulfur cluster) operon of Mycobacterium tuberculosis lacks several genes known to be essential for the function of this system in other organisms. However, the cysteine desulfurase IscSMtb (Rv number Rv3025c; Mtb denotes M. tuberculosis) is conserved in this important pathogen. The present study demonstrates that deleting iscSMtb renders the cells microaerophilic and hypersensitive to oxidative stress. Moreover, the ∆iscSMtb mutant shows impaired Fe-S cluster-dependent enzyme activity, clearly indicating that IscSMtb is associated with Fe-S cluster assembly. An extensive interaction network of IscSMtb with Fe-S proteins was identified, suggesting a novel mechanism of sulfur transfer by direct interaction with apoproteins. Interestingly, the highly homologous IscS of Escherichia coli failed to complement the ∆iscSMtb mutant and showed a less diverse protein-interaction profile. To identify a structural basis for these observations we determined the crystal structure of IscSMtb, which mirrors adaptations made in response to an ISC operon devoid of IscU-like Fe-S cluster scaffold proteins. We conclude that in M. tuberculosis IscS has been redesigned during evolution to compensate for the deletion of large parts of the ISC operon.
- *Global Health Institute, Ecole Polytechnique Fédérale de Lausanne (EPFL), CH-1015 Lausanne, Switzerland.
Organizational Affiliation: