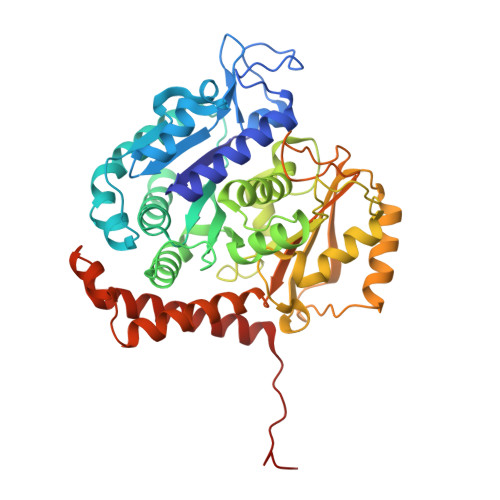
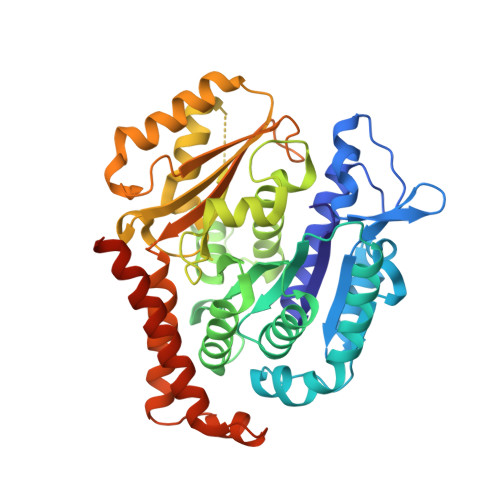
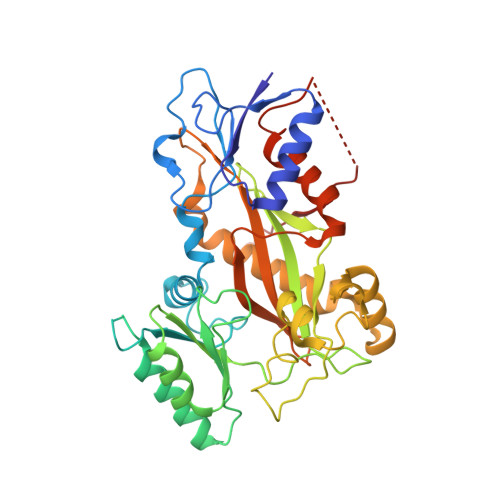
Molecular Mechanism of Action of Microtubule-Stabilizing Anticancer Agents.
Prota, A.E., Bargsten, K., Zurwerra, D., Field, J.J., Diaz, J.F., Altmann, K.H., Steinmetz, M.O.(2013) Science 339: 587-590
- PubMed: 23287720 Search on PubMed
- DOI: https://doi.org/10.1126/science.1230582
- Primary Citation Related Structures:
4I4T, 4I50, 4I55 - PubMed Abstract:
Microtubule-stabilizing agents (MSAs) are efficacious chemotherapeutic drugs widely used for the treatment of cancer. Despite the importance of MSAs for medical applications and basic research, their molecular mechanisms of action on tubulin and microtubules remain elusive. We determined high-resolution crystal structures of αβ-tubulin in complex with two unrelated MSAs, zampanolide and epothilone A. Both compounds were bound to the taxane pocket of β-tubulin and used their respective side chains to induce structuring of the M-loop into a short helix. Because the M-loop establishes lateral tubulin contacts in microtubules, these findings explain how taxane-site MSAs promote microtubule assembly and stability. Further, our results offer fundamental structural insights into the control mechanisms of microtubule dynamics.
- Biomolecular Research, Paul Scherrer Institut, Villigen PSI, Switzerland.
Organizational Affiliation: