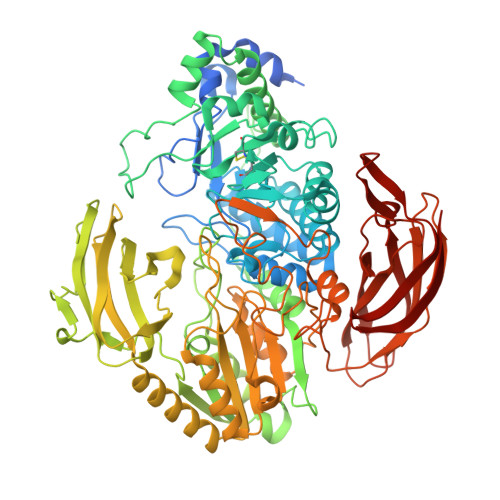
The structure of DesR from Streptomyces venezuelae, a beta-glucosidase involved in macrolide activation.
Zmudka, M.W., Thoden, J.B., Holden, H.M.(2013) Protein Sci 22: 883-892
- PubMed: 23225731
- DOI: https://doi.org/10.1002/pro.2204
- Primary Citation of Related Structures:
4I3G - PubMed Abstract:
Antibiotics have, indeed, altered the course of human history as is evidenced by the increase in human life expectancy since the 1940s. Many of these natural compounds are produced by bacteria that, by necessity, must have efficient self-resistance mechanisms. The methymycin/pikromycin producing species Streptomyces venezuelae, for example, utilizes β-glucosylation of its macrolide products to neutralize their effects within the confines of the cell. Once released into the environment, these compounds are activated by the removal of the glucose moiety. In S. venezuelae, the enzyme responsible for removal of the sugar from the parent compound is encoded by the desR gene and referred to as DesR. It is a secreted enzyme containing 828 amino acid residues, and it is known to be a retaining glycosidase. Here, we describe the structure of the DesR/D-glucose complex determined to 1.4-Å resolution. The overall architecture of the enzyme can be envisioned in terms of three regions: a catalytic core and two auxiliary domains. The catalytic core harbors the binding platform for the glucose ligand. The first auxiliary domain adopts a "PA14 fold," whereas the second auxiliary domain contains an immunoglobulin-like fold. Asp 273 and Glu 578 are in the proper orientation to function as the catalytic base and proton donor, respectively, required for catalysis. The overall fold of the core region places DesR into the GH3 glycoside hydrolase family of enzymes. Comparison of the DesR structure with the β-glucosidase from Kluyveromyces marxianus shows that their PA14 domains assume remarkably different orientations.
- Department of Biochemistry, University of Wisconsin, Madison, Wisconsin 53706, USA.
Organizational Affiliation: