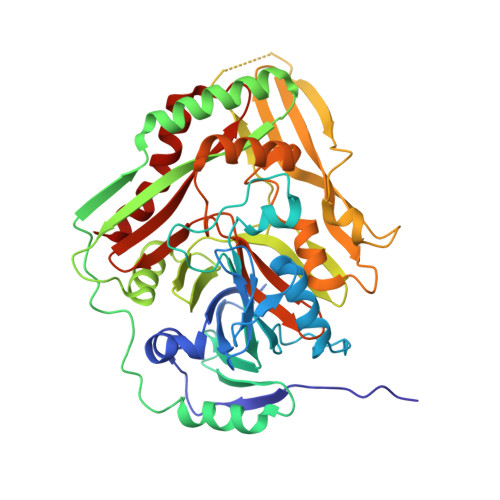
Structure of Aminodeoxychorismate Synthase from Stenotrophomonas maltophilia.
Bera, A.K., Atanasova, V., Dhanda, A., Ladner, J.E., Parsons, J.F.(2012) Biochemistry 51: 10208-10217
- PubMed: 23230967
- DOI: https://doi.org/10.1021/bi301243v
- Primary Citation Related Structures:
4GRH - PubMed Abstract:
PabB, aminodeoxychorismate synthase, is the chorismic acid binding component of the heterodimeric PabA-PabB complex that converts chorismic acid to 4-amino-4-deoxychorismate, a precursor of p-aminobenzoate and folic acid in microorganisms. The second component, a glutamine amidotransferase subunit, PabA, generates ammonia that is channeled to the PabB active site where it attacks C4 of a chorismate-derived intermediate that is covalently bound, through C2, to an active site lysine residue. The presence of a PIKGT motif was, until recently, believed to allow discrimination of PabB enzymes from the closely related enzyme anthranilate synthase, which typically contains a PIAGT active site motif and does not form a covalent enzyme-substrate intermediate with chorismate. A subclass of PabB enzymes that employ an alternative mechanism requiring 2 equiv of ammonia from glutamine and that feature a noncovalently bound 2-amino-2-deoxyisochorismate intermediate was recently identified. Here we report the 2.25 Å crystal structure of PabB from the emerging pathogen Stenotrophomonas maltophilia. It is the first reported structure of a PabB that features the PIAGT motif. Surprisingly, no dedicated pabA is evident in the genome of S. maltophilia, suggesting that another cellular amidotransferase is able to fulfill the role of PabA in this organism. Evaluation of the ammonia-dependent aminodeoxychorismate synthase activity of S. maltophilia PabB alone revealed that it is virtually inactive. However, in the presence of a heterologous PabA surrogate, typical levels of activity were observed using either glutamine or ammonia as the nitrogen source. Additionally, the structure suggests that a key segment of the polypeptide can remodel itself to interact with a nonspecialized or shared amidotransferase partner in vivo. The structure and mass spectral analysis further suggest that S. maltophilia PabB, like Escherichia coli PabB, binds tryptophan in a vestigial regulatory site. The observation that the binding site is unoccupied in the crystal structure, however, suggests the affinity may be low relative to that of E. coli PabB.
- Institute for Bioscience and Biotechnology Research, University of Maryland, MD, USA.
Organizational Affiliation: