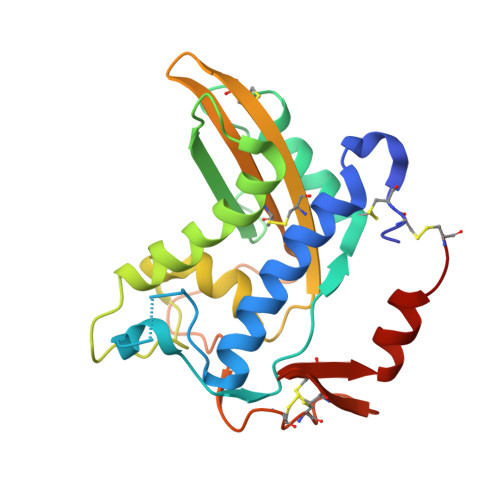
Structure of Ostertagia ostertagi ASP-1: insights into disulfide-mediated cyclization and dimerization
Borloo, J., Geldhof, P., Peelaers, I., Van Meulder, F., Ameloot, P., Callewaert, N., Vercruysse, J., Claerebout, E., Strelkov, S.V., Weeks, S.D.(2013) Acta Crystallogr D Biol Crystallogr 69: 493-503
- PubMed: 23519657
- DOI: https://doi.org/10.1107/S0907444912050019
- Primary Citation Related Structures:
4G2U - PubMed Abstract:
The cysteine-rich secretory/antigen 5/pathogenesis-related 1 (CAP) protein superfamily is composed of a functionally diverse group of members that are found in both eukaryotes and prokaryotes. The excretome/secretome of numerous helminths (parasitic nematodes) contains abundant amounts of CAP members termed activation-associated secreted proteins (ASPs). Although ASPs are necessary for the parasitic life cycle in the host, the current lack of structural and functional information limits both understanding of their actual role in host-parasite interactions and the development of new routes in controlling parasitic infections and diseases. Alleviating this knowledge gap, a 1.85 Å resolution structure of recombinantly produced Oo-ASP-1 from Ostertagia ostertagi, which is one of the most prevalent gastrointestinal parasites in cattle worldwide, was solved. Overall, Oo-ASP-1 displays the common hallmark architecture shared by all CAP-superfamily members, including the N-terminal CAP and C-terminal cysteine-rich domains, but it also reveals a number of highly peculiar features. In agreement with studies of the natively produced protein, the crystal structure shows that Oo-ASP-1 forms a stable dimer that has been found to be primarily maintained via an intermolecular disulfide bridge, hence the small interaction surface of only 306.8 Å(2). Moreover, unlike any other ASP described to date, an additional intramolecular disulfide bridge links the N- and C-termini of each monomer, thereby yielding a quasi-cyclic molecule. Taken together, the insights presented here form an initial step towards a better understanding of the actual biological role(s) that this ASP plays in host-parasite interactions. The structure is also essential to help to define the key regions of the protein suitable for development of ASP-based vaccines, which would enable the current issues surrounding anthelmintic resistance in the treatment of parasitic infections and diseases to be circumvented.
- Laboratory of Parasitology, Faculty of Veterinary Medicine, Ghent University, Merelbeke, Belgium. jimmy.borloo@ugent.be
Organizational Affiliation: