Acyl Guanidine Inhibitors of beta-Secretase (BACE-1): Optimization of a Micromolar Hit to a Nanomolar Lead via Iterative Solid- and Solution-Phase Library Synthesis
Gerritz, S.W., Zhai, W., Shi, S., Zhu, S., Toyn, J.H., Meredith, J.E., Iben, L.G., Burton, C.R., Albright, C.F., Good, A.C., Tebben, A.J., Muckelbauer, J.K., Camac, D.M., Metzler, W., Cook, L.S., Padmanabha, R., Lentz, K.A., Sofia, M.J., Poss, M.A., Macor, J.E., Thompson, L.A.(2012) J Med Chem 55: 9208-9223
- PubMed: 23030502 Search on PubMed
- DOI: https://doi.org/10.1021/jm300931y
- Primary Citation Related Structures:
4FSE, 4FSL - PubMed Abstract:
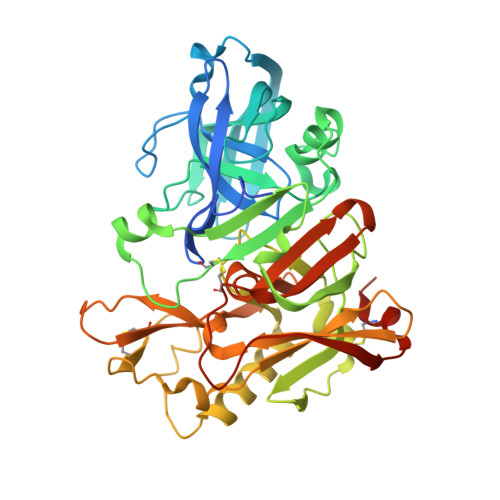
This report describes the discovery and optimization of a BACE-1 inhibitor series containing an unusual acyl guanidine chemotype that was originally synthesized as part of a 6041-membered solid-phase library. The synthesis of multiple follow-up solid- and solution-phase libraries facilitated the optimization of the original micromolar hit into a single-digit nanomolar BACE-1 inhibitor in both radioligand binding and cell-based functional assay formats. The X-ray structure of representative inhibitors bound to BACE-1 revealed a number of key ligand:protein interactions, including a hydrogen bond between the side chain amide of flap residue Gln73 and the acyl guanidine carbonyl group, and a cation-π interaction between Arg235 and the isothiazole 4-methoxyphenyl substituent. Following subcutaneous administration in rats, an acyl guanidine inhibitor with single-digit nanomolar activity in cells afforded good plasma exposures and a dose-dependent reduction in plasma Aβ levels, but poor brain exposure was observed (likely due to Pgp-mediated efflux), and significant reductions in brain Aβ levels were not obtained.
- Bristol-Myers Squibb Research, 5 Research Parkway, Wallingford, Connecticut 06492, United States. samuel.gerritz@bms.com
Organizational Affiliation: