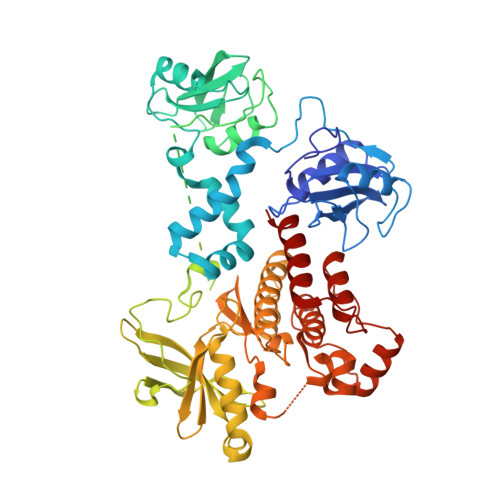
Structural and biophysical characterization of the syk activation switch.
Gradler, U., Schwarz, D., Dresing, V., Musil, D., Bomke, J., Frech, M., Greiner, H., Jakel, S., Rysiok, T., Muller-Pompalla, D., Wegener, A.(2013) J Mol Biology 425: 309-333
- PubMed: 23154170 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2012.11.007
- Primary Citation Related Structures:
4FL1, 4FL2, 4FL3 - PubMed Abstract:
Syk is an essential non-receptor tyrosine kinase in intracellular immunological signaling, and the control of Syk kinase function is considered as a valuable target for pharmacological intervention in autoimmune or inflammation diseases. Upon immune receptor stimulation, the kinase activity of Syk is regulated by binding of phosphorylated immune receptor tyrosine-based activating motifs (pITAMs) to the N-terminal tandem Src homology 2 (tSH2) domain and by autophosphorylation with consequences for the molecular structure of the Syk protein. Here, we present the first crystal structures of full-length Syk (fl-Syk) as wild type and as Y348F,Y352F mutant forms in complex with AMP-PNP revealing an autoinhibited conformation. The comparison with the crystal structure of the truncated Syk kinase domain in complex with AMP-PNP taken together with ligand binding studies by surface plasmon resonance (SPR) suggests conformational differences in the ATP sites of autoinhibited and activated Syk forms. This hypothesis was corroborated by studying the thermodynamic and kinetic interaction of three published Syk inhibitors with isothermal titration calorimetry and SPR, respectively. We further demonstrate the modulation of inhibitor binding affinities in the presence of pITAM and discuss the observed differences of thermodynamic and kinetic signatures. The functional relevance of pITAM binding to fl-Syk was confirmed by a strong stimulation of in vitro autophosphorylation. A structural feedback mechanism on the kinase domain upon pITAM binding to the tSH2 domain is discussed in analogy of the related family kinase ZAP-70 (Zeta-chain-associated protein kinase 70). Surprisingly, we observed distinct conformations of the tSH2 domain and the activation switch including Tyr348 and Tyr352 in the interdomain linker of Syk in comparison to ZAP-70.
- Merck KGaA, Merck Serono Research, Small Molecule Platform/MIB, Frankfurter Str. 250, 64293 Darmstadt, Germany. ulrich.graedler@merckgroup.com
Organizational Affiliation: