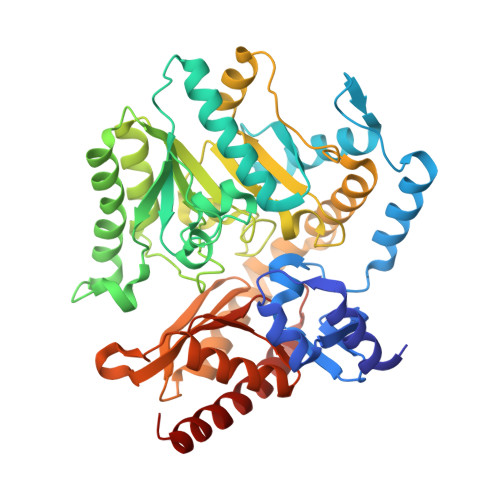
Crystal Structures of the Chromobacterium Violaceum Omega-Transaminase Reveal Major Structural Rearrangements Upon Binding of Coenzyme Plp.
Svedendahl Humble, M., Engelmark Cassimjee, K., Hakansson, M., Kimbung, Y.R., Walse, B., Abedi, V., Federsel, H.-J., Berglund, P., Logan, D.T.(2012) FEBS J 279: 779
- PubMed: 22268978
- DOI: https://doi.org/10.1111/j.1742-4658.2012.08468.x
- Primary Citation Related Structures:
4A6R, 4A6T, 4A6U, 4A72 - PubMed Abstract:
The bacterial ω-transaminase from Chromobacterium violaceum (Cv-ωTA, EC2.6.1.18) catalyses industrially important transamination reactions by use of the coenzyme pyridoxal 5'-phosphate (PLP). Here, we present four crystal structures of Cv-ωTA: two in the apo form, one in the holo form and one in an intermediate state, at resolutions between 1.35 and 2.4 Å. The enzyme is a homodimer with a molecular mass of ∼ 100 kDa. Each monomer has an active site at the dimeric interface that involves amino acid residues from both subunits. The apo-Cv-ωTA structure reveals unique 'relaxed' conformations of three critical loops involved in structuring the active site that have not previously been seen in a transaminase. Analysis of the four crystal structures reveals major structural rearrangements involving elements of the large and small domains of both monomers that reorganize the active site in the presence of PLP. The conformational change appears to be triggered by binding of the phosphate group of PLP. Furthermore, one of the apo structures shows a disordered 'roof ' over the PLP-binding site, whereas in the other apo form and the holo form the 'roof' is ordered. Comparison with other known transaminase crystal structures suggests that ordering of the 'roof' structure may be associated with substrate binding in Cv-ωTA and some other transaminases. The atomic coordinates and structure factors for the Chromobacterium violaceumω-transaminase crystal structures can be found in the RCSB Protein Data Bank (http://www.rcsb.org) under the accession codes 4A6U for the holoenzyme, 4A6R for the apo1 form, 4A6T for the apo2 form and 4A72 for the mixed form • -transaminases and -transaminases bind by dynamic light scattering (View interaction) • -transaminase and -transaminase bind by x-ray crystallography (View interaction) • -transaminase and -transaminase bind by x-ray crystallography (View interaction).
- Division of Biochemistry, KTH Royal Institute of Technology, Stockholm, Sweden.
Organizational Affiliation: