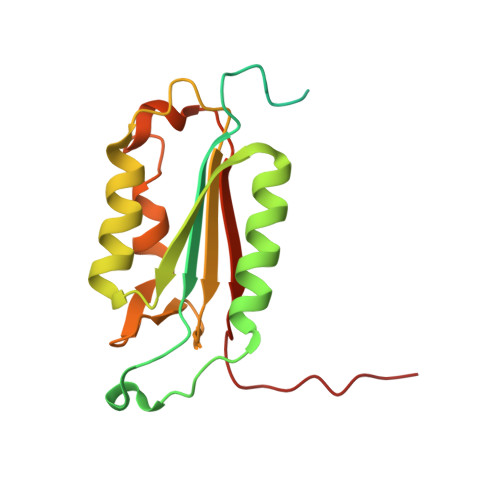
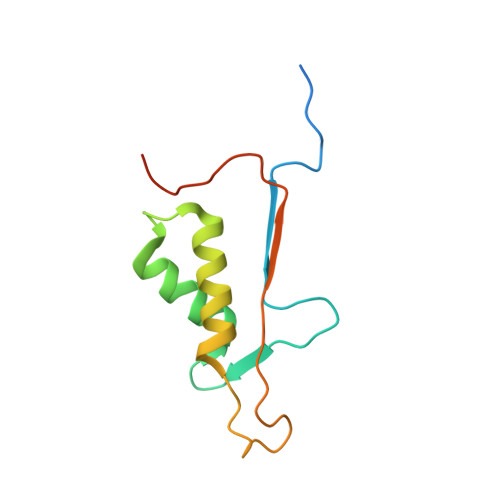
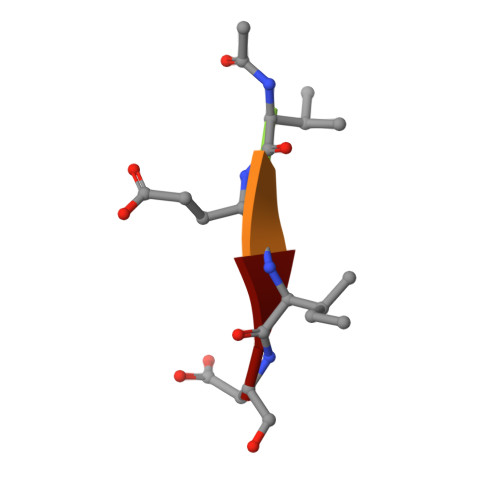
Reprogramming Caspase-7 Specificity by Regio-Specific Mutations and Selection Provides Alternate Solutions for Substrate Recognition.
Hill, M.E., MacPherson, D.J., Wu, P., Julien, O., Wells, J.A., Hardy, J.A.(2016) ACS Chem Biol 11: 1603-1612
- PubMed: 27032039 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acschembio.5b00971
- Primary Citation Related Structures:
4ZVO, 4ZVP, 4ZVQ, 4ZVR, 4ZVS, 4ZVT, 4ZVU - PubMed Abstract:
The ability to routinely engineer protease specificity can allow us to better understand and modulate their biology for expanded therapeutic and industrial applications. Here, we report a new approach based on a caged green fluorescent protein (CA-GFP) reporter that allows for flow-cytometry-based selection in bacteria or other cell types enabling selection of intracellular protease specificity, regardless of the compositional complexity of the protease. Here, we apply this approach to introduce the specificity of caspase-6 into caspase-7, an intracellular cysteine protease important in cellular remodeling and cell death. We found that substitution of substrate-contacting residues from caspase-6 into caspase-7 was ineffective, yielding an inactive enzyme, whereas saturation mutagenesis at these positions and selection by directed evolution produced active caspases. The process produced a number of nonobvious mutations that enabled conversion of the caspase-7 specificity to match caspase-6. The structures of the evolved-specificity caspase-7 (esCasp-7) revealed alternate binding modes for the substrate, including reorganization of an active site loop. Profiling the entire human proteome of esCasp-7 by N-terminomics demonstrated that the global specificity toward natural protein substrates is remarkably similar to that of caspase-6. Because the esCasp-7 maintained the core of caspase-7, we were able to identify a caspase-6 substrate, lamin C, that we predict relies on an exosite for substrate recognition. These reprogrammed proteases may be the first tool built with the express intent of distinguishing exosite dependent or independent substrates. This approach to specificity reprogramming should also be generalizable across a wide range of proteases.
- Department of Chemistry, 104 LGRT, 710 N. Pleasant St., University of Massachusetts , Amherst, Massachusetts 01003, United States.
Organizational Affiliation: