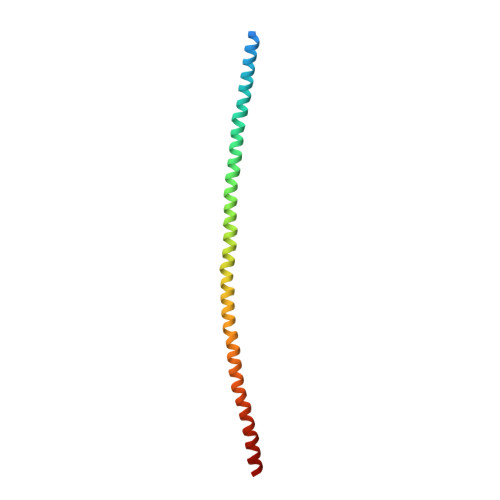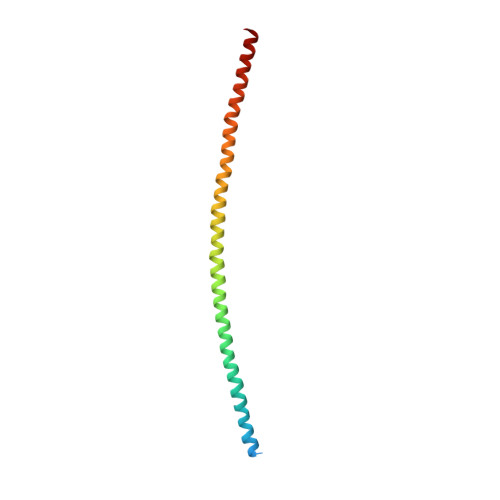The X-Ray Crystal Structure of the Keratin 1-Keratin 10 Helix 2B Heterodimer Reveals Molecular Surface Properties and Biochemical Insights into Human Skin Disease.
Bunick, C.G., Milstone, L.M.(2017) J Invest Dermatol 137: 142-150
- PubMed: 27595935 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jid.2016.08.018
- Primary Citation Related Structures:
4ZRY - PubMed Abstract:
Keratins 1 (K1) and 10 (K10) are the primary keratins expressed in differentiated epidermis. Mutations in K1/K10 are associated with human skin diseases. We determined the crystal structure of the complex between the distal (2B) helices of K1 and K10 to better understand how human keratin structure correlates with function. The 3.3 Å resolution structure confirms many features inferred by previous biochemical analyses, but adds unexpected insights. It demonstrates a parallel, coiled-coil heterodimer with a predominantly hydrophobic intermolecular interface; this heterodimer formed a higher order complex with a second K1-K10-2B heterodimer via a Cys401 K10 disulfide link, although the bond angle is unanticipated. The molecular surface analysis of K1-K10-2B identified several pockets, one adjacent to the disulfide linkage and conserved in K5-K14. The solvent accessible surface area of the K1-K10 structure is 20-25% hydrophobic. The 2B region contains mixed acidic and basic patches proximally (N-terminal), whereas it is largely acidic distally (C-terminal). Mapping of conserved and nonconserved residues between K1-K10 and K5-K14 onto the structure demonstrated the majority of unique residues align along the outer helical ridge. Finally, the structure permitted a fresh analysis of the deleterious effects caused by K1/K10 missense mutations found in patients with phenotypic skin disease.
- Department of Dermatology, Yale University, New Haven, Connecticut, USA. Electronic address: christopher.bunick@yale.edu.
Organizational Affiliation: