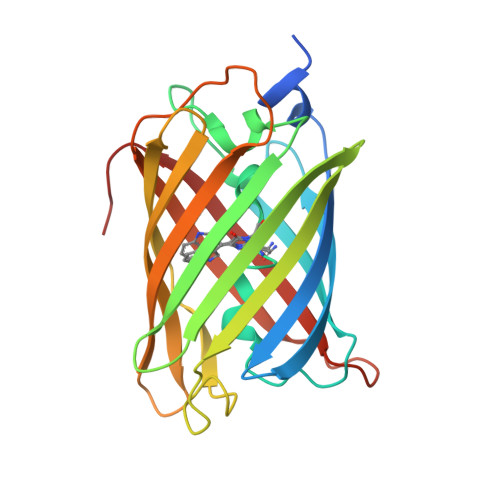
Crystal Structure of Phototoxic Orange Fluorescent Proteins with a Tryptophan-Based Chromophore.
Pletneva, N.V., Pletnev, V.Z., Sarkisyan, K.S., Gorbachev, D.A., Egorov, E.S., Mishin, A.S., Lukyanov, K.A., Dauter, Z., Pletnev, S.(2015) PLoS One 10: e0145740-e0145740
- PubMed: 26699366 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0145740
- Primary Citation Related Structures:
4ZBL, 4ZFS - PubMed Abstract:
Phototoxic fluorescent proteins represent a sparse group of genetically encoded photosensitizers that could be used for precise light-induced inactivation of target proteins, DNA damage, and cell killing. Only two such GFP-based fluorescent proteins (FPs), KillerRed and its monomeric variant SuperNova, were described up to date. Here, we present a crystallographic study of their two orange successors, dimeric KillerOrange and monomeric mKillerOrange, at 1.81 and 1.57 Å resolution, respectively. They are the first orange-emitting protein photosensitizers with a tryptophan-based chromophore (Gln65-Trp66-Gly67). Same as their red progenitors, both orange photosensitizers have a water-filled channel connecting the chromophore to the β-barrel exterior and enabling transport of ROS. In both proteins, Trp66 of the chromophore adopts an unusual trans-cis conformation stabilized by H-bond with the nearby Gln159. This trans-cis conformation along with the water channel was shown to be a key structural feature providing bright orange emission and phototoxicity of both examined orange photosensitizers.
- Shemyakin-Ovchinnikov Institute of Bioorganic Chemistry, Russian Academy of Sciences, Moscow, Russian Federation.
Organizational Affiliation: