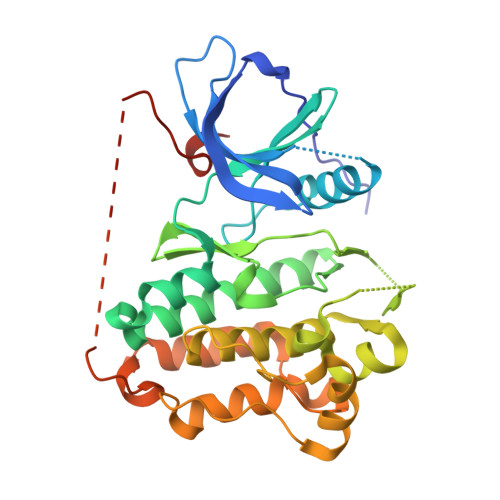
Binding mode of the breakthrough inhibitor AZD9291 to epidermal growth factor receptor revealed.
Yosaatmadja, Y., Silva, S., Dickson, J.M., Patterson, A.V., Smaill, J.B., Flanagan, J.U., McKeage, M.J., Squire, C.J.(2015) J Struct Biol 192: 539-544
- PubMed: 26522274 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2015.10.018
- Primary Citation Related Structures:
4ZAU - PubMed Abstract:
The discovery of genetic drivers of lung cancer in patient sub-groups has led to their use as predictive biomarkers and as targets for selective drug therapy. Some of the most important lung cancer drivers are mutations in the EGFR gene, for example, the exon 19 deletions and the L858R variant that confer sensitivity to the front line drugs erlotinib and gefitinib; the acquired T790M variants confer drug resistance and a poor prognosis. A challenge then in targeting EGFR is to produce drugs that inhibit both sensitising variants and resistance variants, leaving wild type protein in healthy cells unaffected. One such agent is AstraZeneca's "breakthrough" AZD9291 molecule that shows a 200-fold selectivity for T790M/L858R over wild type EGFR. Our X-ray crystal structure reveals the binding mode of AZD9291 to the kinase domain of wild type EGFR.
- School of Biological Sciences, The University of Auckland, Private Bag 92019, Auckland, New Zealand.
Organizational Affiliation: