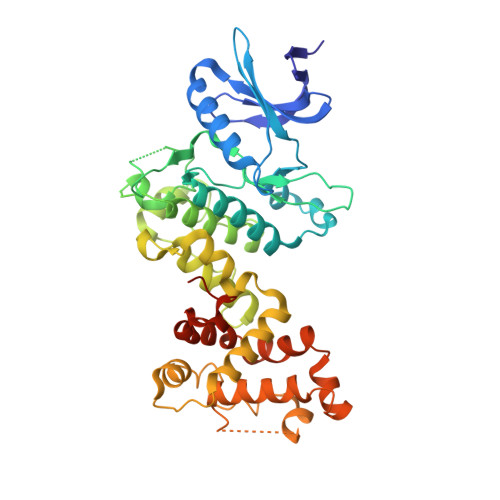
Molecular mechanisms of human IRE1 activation through dimerization and ligand binding.
Joshi, A., Newbatt, Y., McAndrew, P.C., Stubbs, M., Burke, R., Richards, M.W., Bhatia, C., Caldwell, J.J., McHardy, T., Collins, I., Bayliss, R.(2015) Oncotarget 6: 13019-13035
- PubMed: 25968568 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.18632/oncotarget.3864
- Primary Citation Related Structures:
4Z7G, 4Z7H - PubMed Abstract:
IRE1 transduces the unfolded protein response by splicing XBP1 through its C-terminal cytoplasmic kinase-RNase region. IRE1 autophosphorylation is coupled to RNase activity through formation of a back-to-back dimer, although the conservation of the underlying molecular mechanism is not clear from existing structures. We have crystallized human IRE1 in a back-to-back conformation only previously seen for the yeast homologue. In our structure the kinase domain appears primed for catalysis but the RNase domains are disengaged. Structure-function analysis reveals that IRE1 is autoinhibited through a Tyr-down mechanism related to that found in the unrelated Ser/Thr protein kinase Nek7. We have developed a compound that potently inhibits human IRE1 kinase activity while stimulating XBP1 splicing. A crystal structure of the inhibitor bound to IRE1 shows an increased ordering of the kinase activation loop. The structures of hIRE in apo and ligand-bound forms are consistent with a previously proposed model of IRE1 regulation in which formation of a back-to-back dimer coupled to adoption of a kinase-active conformation drive RNase activation. The structures provide opportunities for structure-guided design of IRE1 inhibitors.
- Department of Biochemistry, University of Leicester, Leicester, United Kingdom.
Organizational Affiliation: