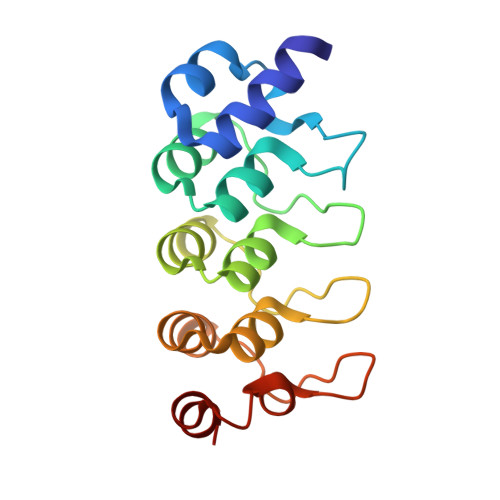
Hybrid Structural Analysis of the Arp2/3 Regulator Arpin Identifies Its Acidic Tail as a Primary Binding Epitope.
Fetics, S., Thureau, A., Campanacci, V., Aumont-Nicaise, M., Dang, I., Gautreau, A., Perez, J., Cherfils, J.(2016) Structure 24: 252-260
- PubMed: 26774128 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2015.12.001
- Primary Citation Related Structures:
4Z68 - PubMed Abstract:
Arpin is a newly discovered regulator of actin polymerization at the cell leading edge, which steers cell migration by exerting a negative control on the Arp2/3 complex. Arpin proteins have an acidic tail homologous to the acidic motif of the VCA domain of nucleation-promoting factors (NPFs). This tail is predicted to compete with the VCA of NPFs for binding to the Arp2/3 complex, thereby mitigating activation and/or tethering of the complex to sites of actin branching. Here, we investigated the structure of full-length Arpin using synchrotron small-angle X-ray scattering, and of its acidic tail in complex with an ankyrin repeats domain using X-ray crystallography. The data were combined in a hybrid model in which the acidic tail extends from the globular core as a linear peptide and forms a primary epitope that is readily accessible in unbound Arpin and suffices to tether Arpin to interacting proteins with high affinity.
- Laboratoire de Pharmacologie et Biologie Appliquée, UMR 8113, CNRS-Ecole Normale Supérieure de Cachan, 61 Avenue du Président Wilson, 94235 Cachan Cedex, France; Laboratoire d'Enzymologie et Biochimie Structurales, CNRS UPR3082, 91190 Gif-sur-Yvette, France.
Organizational Affiliation: