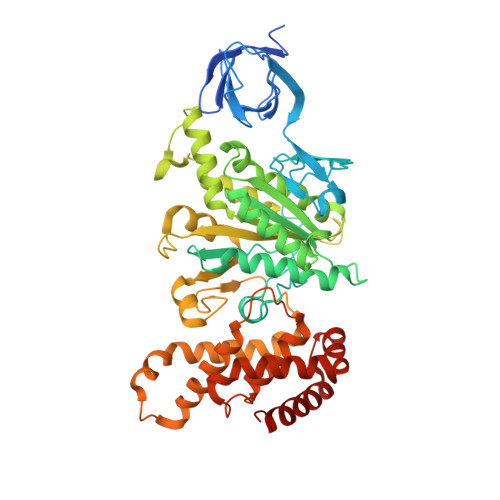
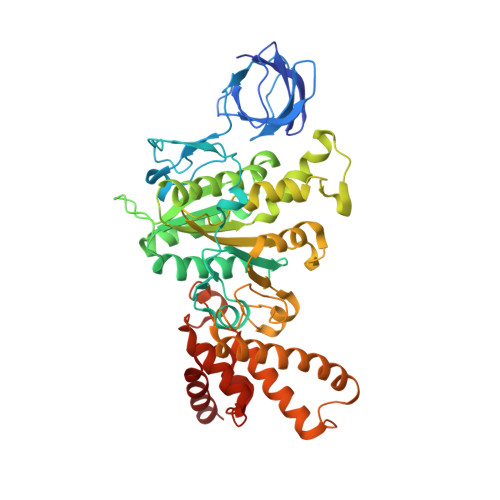
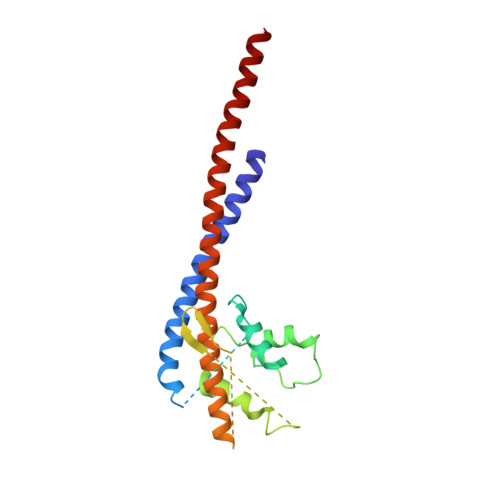
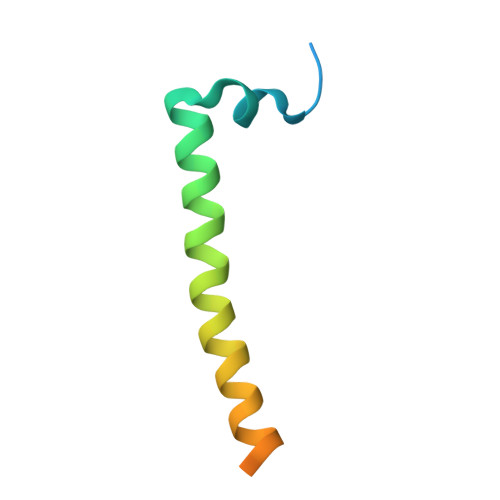
How release of phosphate from mammalian F1-ATPase generates a rotary substep.
Bason, J.V., Montgomery, M.G., Leslie, A.G., Walker, J.E.(2015) Proc Natl Acad Sci U S A 112: 6009-6014
- PubMed: 25918412 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1506465112
- Primary Citation Related Structures:
4YXW, 4Z1M - PubMed Abstract:
The rotation of the central stalk of F1-ATPase is driven by energy derived from the sequential binding of an ATP molecule to its three catalytic sites and the release of the products of hydrolysis. In human F1-ATPase, each 360° rotation consists of three 120° steps composed of substeps of about 65°, 25°, and 30°, with intervening ATP binding, phosphate release, and catalytic dwells, respectively. The F1-ATPase inhibitor protein, IF1, halts the rotary cycle at the catalytic dwell. The human and bovine enzymes are essentially identical, and the structure of bovine F1-ATPase inhibited by IF1 represents the catalytic dwell state. Another structure, described here, of bovine F1-ATPase inhibited by an ATP analog and the phosphate analog, thiophosphate, represents the phosphate binding dwell. Thiophosphate is bound to a site in the α(E)β(E)-catalytic interface, whereas in F1-ATPase inhibited with IF1, the equivalent site is changed subtly and the enzyme is incapable of binding thiophosphate. These two structures provide a molecular mechanism of how phosphate release generates a rotary substep as follows. In the active enzyme, phosphate release from the β(E)-subunit is accompanied by a rearrangement of the structure of its binding site that prevents released phosphate from rebinding. The associated extrusion of a loop in the β(E)-subunit disrupts interactions in the α(E)β(E-)catalytic interface and opens it to its fullest extent. Other rearrangements disrupt interactions between the γ-subunit and the C-terminal domain of the α(E)-subunit. To restore most of these interactions, and to make compensatory new ones, the γ-subunit rotates through 25°-30°.
- Medical Research Council Mitochondrial Biology Unit, Cambridge Biomedical Campus, Hills Road, Cambridge CB2 0XY, United Kingdom; and.
Organizational Affiliation: