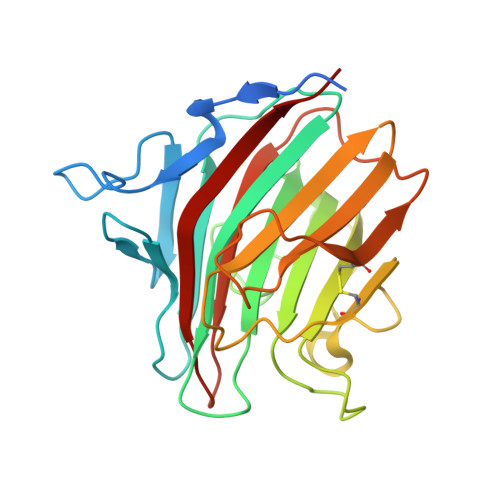
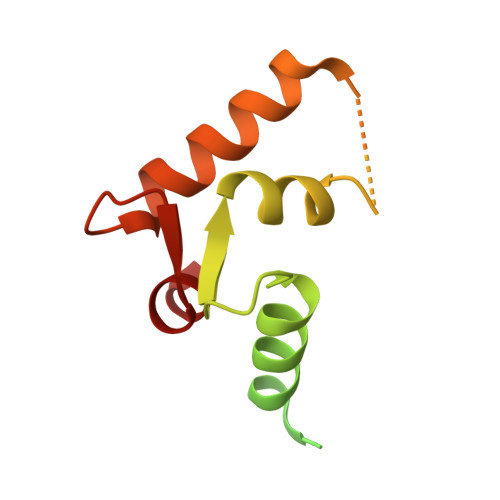
Crystallographic snapshots of the EF-hand protein MCFD2 complexed with the intracellular lectin ERGIC-53 involved in glycoprotein transport.
Satoh, T., Nishio, M., Suzuki, K., Yagi-Utsumi, M., Kamiya, Y., Mizushima, T., Kato, K.(2020) Acta Crystallogr F Struct Biol Commun 76: 216-221
- PubMed: 32356523 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X20005452
- Primary Citation Related Structures:
4YGB, 4YGC, 4YGD, 4YGE - PubMed Abstract:
The transmembrane intracellular lectin ER-Golgi intermediate compartment protein 53 (ERGIC-53) and the soluble EF-hand multiple coagulation factor deficiency protein 2 (MCFD2) form a complex that functions as a cargo receptor, trafficking various glycoproteins between the endoplasmic reticulum (ER) and the Golgi apparatus. It has been demonstrated that the carbohydrate-recognition domain (CRD) of ERGIC-53 (ERGIC-53 CRD ) interacts with N-linked glycans on cargo glycoproteins, whereas MCFD2 recognizes polypeptide segments of cargo glycoproteins. Crystal structures of ERGIC-53 CRD complexed with MCFD2 and mannosyl oligosaccharides have revealed protein-protein and protein-sugar binding modes. In contrast, the polypeptide-recognition mechanism of MCFD2 remains largely unknown. Here, a 1.60 Å resolution crystal structure of the ERGIC-53 CRD -MCFD2 complex is reported, along with three other crystal forms. Comparison of these structures with those previously reported reveal that MCFD2, but not ERGIC-53-CRD, exhibits significant conformational plasticity that may be relevant to its accommodation of various polypeptide ligands.
- Graduate School of Pharmaceutical Sciences, Nagoya City University, 3-1 Tanabe-dori, Mizuho-ku, Nagoya, Aichi 467-8603, Japan.
Organizational Affiliation: