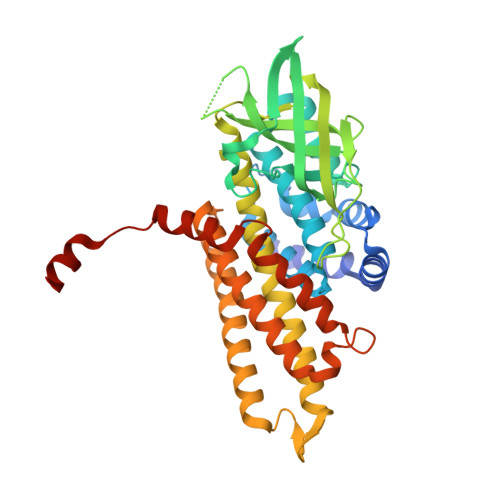
A covalent adduct of MbtN, an acyl-ACP dehydrogenase from Mycobacterium tuberculosis, reveals an unusual acyl-binding pocket.
Chai, A.F., Bulloch, E.M., Evans, G.L., Lott, J.S., Baker, E.N., Johnston, J.M.(2015) Acta Crystallogr D Biol Crystallogr 71: 862-872
- PubMed: 25849397 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004715001650
- Primary Citation Related Structures:
4XVX - PubMed Abstract:
Mycobacterium tuberculosis (Mtb) is the causative agent of tuberculosis. Access to iron in host macrophages depends on iron-chelating siderophores called mycobactins and is strongly correlated with Mtb virulence. Here, the crystal structure of an Mtb enzyme involved in mycobactin biosynthesis, MbtN, in complex with its FAD cofactor is presented at 2.30 Å resolution. The polypeptide fold of MbtN conforms to that of the acyl-CoA dehydrogenase (ACAD) family, consistent with its predicted role of introducing a double bond into the acyl chain of mycobactin. Structural comparisons and the presence of an acyl carrier protein, MbtL, in the same gene locus suggest that MbtN acts on an acyl-(acyl carrier protein) rather than an acyl-CoA. A notable feature of the crystal structure is the tubular density projecting from N(5) of FAD. This was interpreted as a covalently bound polyethylene glycol (PEG) fragment and resides in a hydrophobic pocket where the substrate acyl group is likely to bind. The pocket could accommodate an acyl chain of 14-21 C atoms, consistent with the expected length of the mycobactin acyl chain. Supporting this, steady-state kinetics show that MbtN has ACAD activity, preferring acyl chains of at least 16 C atoms. The acyl-binding pocket adopts a different orientation (relative to the FAD) to other structurally characterized ACADs. This difference may be correlated with the apparent ability of MbtN to catalyse the formation of an unusual cis double bond in the mycobactin acyl chain.
- Laboratory of Structural Biology, School of Biological Sciences and Maurice Wilkins Centre for Molecular Biodiscovery, University of Auckland, Private Bag 92019, Auckland 1142, New Zealand.
Organizational Affiliation: