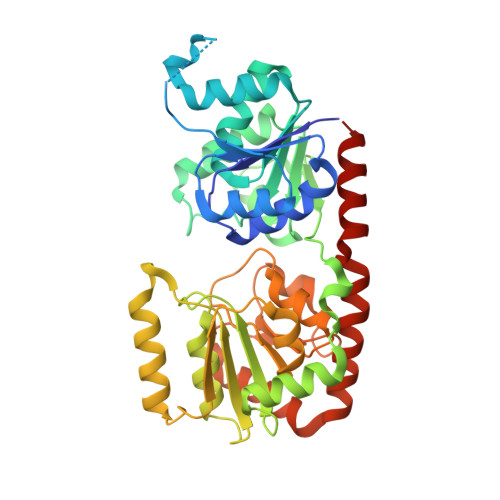
Structural and enzymatic analyses of a glucosyltransferase Alr3699/HepE involved in Anabaena heterocyst envelop polysaccharide biosynthesis
Wang, X.P., Jiang, Y.L., Dai, Y.N., Cheng, W., Chen, Y., Zhou, C.Z.(2016) Glycobiology 26: 520-531
- PubMed: 26692049 Search on PubMed
- DOI: https://doi.org/10.1093/glycob/cwv167
- Primary Citation Related Structures:
4XSO, 4XSP, 4XSR, 4XSU - PubMed Abstract:
Formation of the heterocyst envelope polysaccharide (HEP) is a key process for cyanobacterial heterocyst differentiation. The maturation of HEP in Anabaena sp. strain PCC 7120 is controlled by a gene cluster termed HEP island in addition to an operon alr3698-alr3699, which encodes two putative proteins termed Alr3698/HepD and Alr3699/HepE. Here we report the crystal structures of HepE in the apo-form and three complex forms that bind to UDP-glucose (UDPG), UDP&glucose, and UDP, respectively. The overall structure of HepE displays a typical GT-B fold of glycosyltransferases, comprising two separate β/α/β Rossmann-fold domains that form an inter-domain substrate-binding crevice. Structural analyses combined with enzymatic assays indicate that HepE is a glucosyltransferase using UDPG as a sugar donor. Further site-directed mutageneses enable us to assign the key residues that stabilize the sugar donor and putative acceptor. Based on the comparative structural analyses, we propose a putative catalytic cycle of HepE, which undergoes "open-closed-open" conformational changes upon binding to the substrates and release of products. These findings provide structural and catalytic insights into the first enzyme involved in the HEP biosynthesis pathway.
- Hefei National Laboratory for Physical Sciences at the Microscale and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui 230027, People's Republic of China.
Organizational Affiliation: