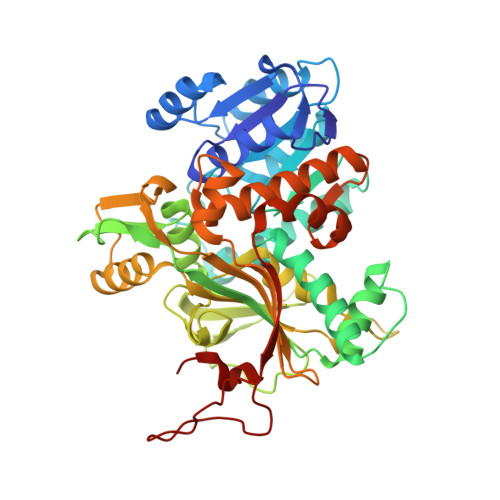
Comprehensive Structural Characterization of the Bacterial Homospermidine Synthase-an Essential Enzyme of the Polyamine Metabolism.
Krossa, S., Faust, A., Ober, D., Scheidig, A.J.(2016) Sci Rep 6: 19501-19501
- PubMed: 26776105
- DOI: https://doi.org/10.1038/srep19501
- Primary Citation of Related Structures:
4PLP, 4TVB, 4XQ9, 4XQC, 4XQE, 4XQG, 4XR4, 4XRG - PubMed Abstract:
The highly conserved bacterial homospermidine synthase (HSS) is a key enzyme of the polyamine metabolism of many proteobacteria including pathogenic strains such as Legionella pneumophila and Pseudomonas aeruginosa; The unique usage of NAD(H) as a prosthetic group is a common feature of bacterial HSS, eukaryotic HSS and deoxyhypusine synthase (DHS). The structure of the bacterial enzyme does not possess a lysine residue in the active center and thus does not form an enzyme-substrate Schiff base intermediate as observed for the DHS. In contrast to the DHS the active site is not formed by the interface of two subunits but resides within one subunit of the bacterial HSS. Crystal structures of Blastochloris viridis HSS (BvHSS) reveal two distinct substrate binding sites, one of which is highly specific for putrescine. BvHSS features a side pocket in the direct vicinity of the active site formed by conserved amino acids and a potential substrate discrimination, guiding, and sensing mechanism. The proposed reaction steps for the catalysis of BvHSS emphasize cation-π interaction through a conserved Trp residue as a key stabilizer of high energetic transition states.
- Structural Biology-Zoological Institute, Kiel University, Am Botanischen Garten 11, 24118 Kiel, Germany.
Organizational Affiliation: