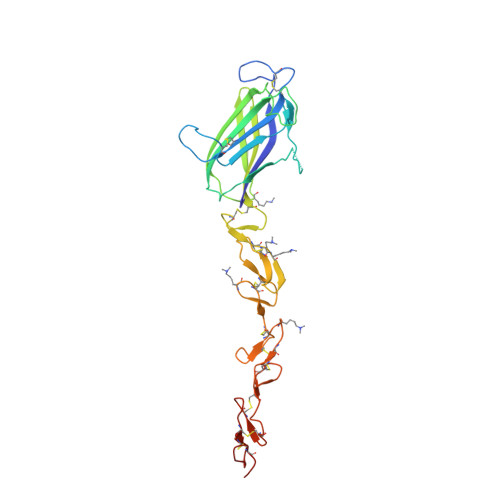
Structural biology. Structural basis for Notch1 engagement of Delta-like 4.
Luca, V.C., Jude, K.M., Pierce, N.W., Nachury, M.V., Fischer, S., Garcia, K.C.(2015) Science 347: 847-853
- PubMed: 25700513 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.1261093
- Primary Citation Related Structures:
4XL1, 4XLW - PubMed Abstract:
Notch receptors guide mammalian cell fate decisions by engaging the proteins Jagged and Delta-like (DLL). The 2.3 angstrom resolution crystal structure of the interacting regions of the Notch1-DLL4 complex reveals a two-site, antiparallel binding orientation assisted by Notch1 O-linked glycosylation. Notch1 epidermal growth factor-like repeats 11 and 12 interact with the DLL4 Delta/Serrate/Lag-2 (DSL) domain and module at the N-terminus of Notch ligands (MNNL) domains, respectively. Threonine and serine residues on Notch1 are functionalized with O-fucose and O-glucose, which act as surrogate amino acids by making specific, and essential, contacts to residues on DLL4. The elucidation of a direct chemical role for O-glycans in Notch1 ligand engagement demonstrates how, by relying on posttranslational modifications of their ligand binding sites, Notch proteins have linked their functional capacity to developmentally regulated biosynthetic pathways.
- Howard Hughes Medical Institute, Stanford University School of Medicine, Stanford, CA 94305, USA. Department of Molecular and Cellular Physiology, Stanford University School of Medicine, Stanford, CA 94305, USA. Department of Structural Biology, Stanford University School of Medicine, Stanford, CA 94305, USA.
Organizational Affiliation: