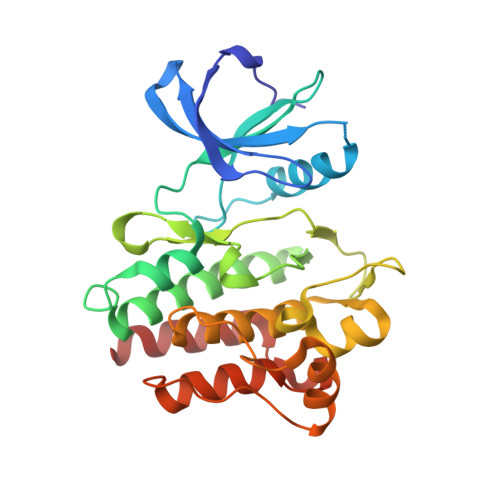
Structure of the ABL2/ARG kinase in complex with dasatinib.
Ha, B.H., Simpson, M.A., Koleske, A.J., Boggon, T.J.(2015) Acta Crystallogr F Struct Biol Commun 71: 443-448
- PubMed: 25849507 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X15004793
- Primary Citation Related Structures:
4XLI - PubMed Abstract:
ABL2/ARG (ABL-related gene) belongs to the ABL (Abelson tyrosine-protein kinase) family of tyrosine kinases. ARG plays important roles in cell morphogenesis, motility, growth and survival, and many of these biological roles overlap with the cellular functions of the ABL kinase. Chronic myeloid leukemia (CML) is associated with constitutive ABL kinase activation resulting from fusion between parts of the breakpoint cluster region (BCR) and ABL1 genes. Similarly, fusion of the ETV6 (Tel) and ARG genes drives some forms of T-cell acute lymphoblastic leukemia (T-ALL) and acute myeloid leukemia (AML). Dasatinib is a tyrosine kinase inhibitor used for the treatment of CML by inhibiting ABL, and while it also inhibits ARG, there is currently no structure of ARG in complex with dasatinib. Here, the co-crystal structure of the mouse ARG catalytic domain with dasatinib at 2.5 Å resolution is reported. Dasatinib-bound ARG is found in the DFG-in conformation although it is nonphosphorylated on the activation-loop tyrosine. In this structure the glycine-rich P-loop is found in a relatively open conformation compared with other known ABL family-inhibitor complex structures.
- Department of Pharmacology, Yale University School of Medicine, 333 Cedar Street, New Haven, CT 06520, USA.
Organizational Affiliation: