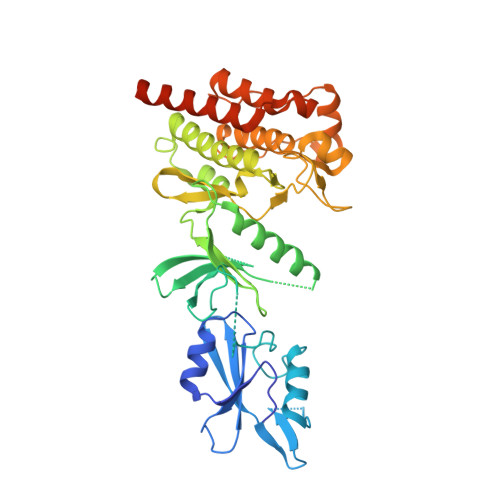
Crystal structure of an SH2-kinase construct of c-Abl and effect of the SH2 domain on kinase activity.
Lorenz, S., Deng, P., Hantschel, O., Superti-Furga, G., Kuriyan, J.(2015) Biochem J 468: 283-291
- PubMed: 25779001 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BJ20141492
- Primary Citation Related Structures:
4XEY - PubMed Abstract:
Constitutive activation of the non-receptor tyrosine kinase c-Abl (cellular Abelson tyrosine protein kinase 1, Abl1) in the Bcr (breakpoint cluster region)-Abl1 fusion oncoprotein is the molecular cause of chronic myeloid leukaemia (CML). Recent studies have indicated that an interaction between the SH2 (Src-homology 2) domain and the N-lobe (N-terminal lobe) of the c-Abl kinase domain (KD) has a critical role in leukaemogenesis [Grebien et al. (2011) Cell 147, 306-319; Sherbenou et al. (2010) Blood 116, 3278-3285]. To dissect the structural basis of this phenomenon, we studied c-Abl constructs comprising the SH2 and KDs in vitro. We present a crystal structure of an SH2-KD construct bound to dasatinib, which contains the relevant interface between the SH2 domain and the N-lobe of the KD. We show that the presence of the SH2 domain enhances kinase activity moderately and that this effect depends on contacts in the SH2/N-lobe interface and is abrogated by specific mutations. Consistently, formation of the interface decreases slightly the association rate of imatinib with the KD. That the effects are small compared with the dramatic in vivo consequences suggests an important function of the SH2-N-lobe interaction might be to help disassemble the auto-inhibited conformation of c-Abl and promote processive phosphorylation, rather than substantially stimulate kinase activity.
- *California Institute for Quantitative Biosciences, University of California at Berkeley, Berkeley, CA 94720, U.S.A.
Organizational Affiliation: