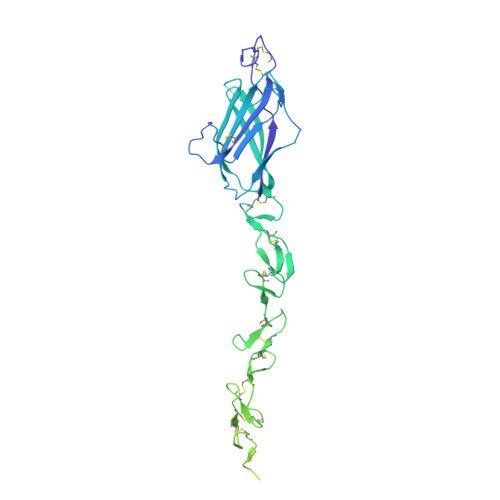
Notch ligand delta-like1: X-ray crystal structure and binding affinity.
Kershaw, N.J., Church, N.L., Griffin, M.D., Luo, C.S., Adams, T.E., Burgess, A.W.(2015) Biochem J 468: 159-166
- PubMed: 25715738 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20150010
- Primary Citation Related Structures:
4XBM - PubMed Abstract:
The Notch pathway is a fundamental signalling system in most multicellular animals. We have determined the X-ray crystal structure of the extracellular domain of the Notch ligand delta-like ligand-1 (Dll-1). The structure incorporates the N-terminal C2 domain, receptor-binding DSL domain and the first six (of eight) EGF (epidermal growth factor)-like repeats, which form a highly extended conformation, confirmed by analytical ultracentrifugation. Comparison of our structure with a fragment of Jagged1 ligand allows us to dissect the similarities and differences between the ligand families. Differences in the C2 domains of Dll-1 and Jagged1 suggest their lipid-binding properties are likely to differ. A conserved hydrophobic patch on the surface of both Dll-1 and Jagged1 provides a likely receptor-interaction site that is common to both ligands. We also explore the binding affinity of Dll-1 for a fragment of Notch1 using different techniques. Apparent binding affinities vary when different techniques are used, explaining discrepancies in the literature. Using analytical ultracentrifugation, we perform for the first time binding analyses where both receptor and ligand are in solution, which confirms a Kd of 10 μM for this interaction.
- *Department of Structural Biology, Walter and Eliza Hall Institute of Medical Research and Department of Medical Biology, University of Melbourne, Parkville, 3052 Australia.
Organizational Affiliation: