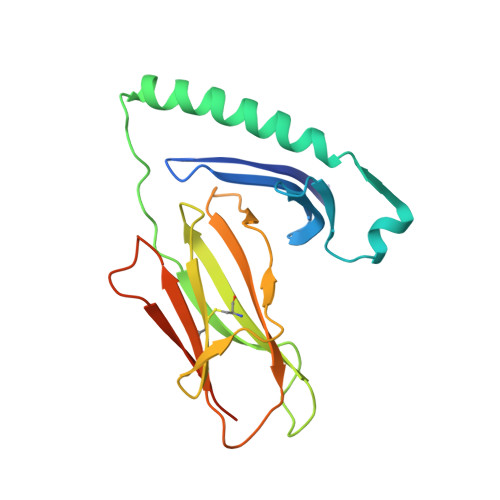
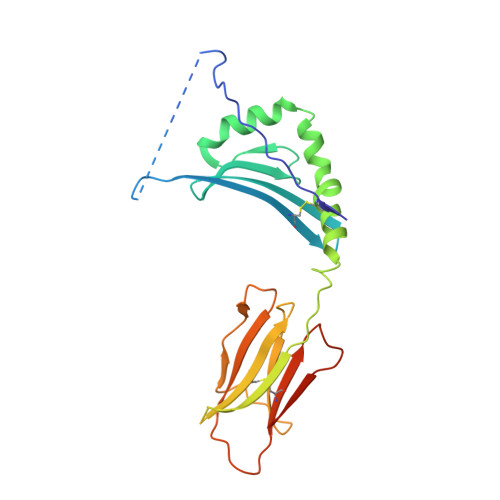
MHC class II complexes sample intermediate states along the peptide exchange pathway.
Wieczorek, M., Sticht, J., Stolzenberg, S., Gunther, S., Wehmeyer, C., El Habre, Z., Alvaro-Benito, M., Noe, F., Freund, C.(2016) Nat Commun 7: 13224-13224
- PubMed: 27827392 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms13224
- Primary Citation Related Structures:
4X5W, 4X5X - PubMed Abstract:
The presentation of peptide-MHCII complexes (pMHCIIs) for surveillance by T cells is a well-known immunological concept in vertebrates, yet the conformational dynamics of antigen exchange remain elusive. By combining NMR-detected H/D exchange with Markov modelling analysis of an aggregate of 275 microseconds molecular dynamics simulations, we reveal that a stable pMHCII spontaneously samples intermediate conformations relevant for peptide exchange. More specifically, we observe two major peptide exchange pathways: the kinetic stability of a pMHCII's ground state defines its propensity for intrinsic peptide exchange, while the population of a rare, intermediate conformation correlates with the propensity of the HLA-DM-catalysed pathway. Helix-destabilizing mutants designed based on our model shift the exchange behaviour towards the HLA-DM-catalysed pathway and further allow us to conceptualize how allelic variation can shape an individual's MHC restricted immune response.
- Protein Biochemistry, Institute for Chemistry and Biochemistry, Freie Universität Berlin, Thielallee 63, 14195 Berlin, Germany.
Organizational Affiliation: