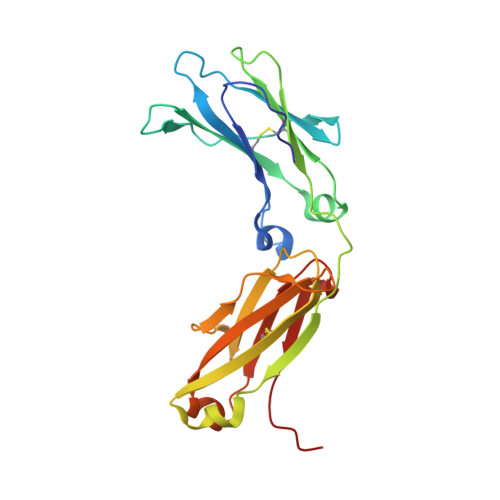
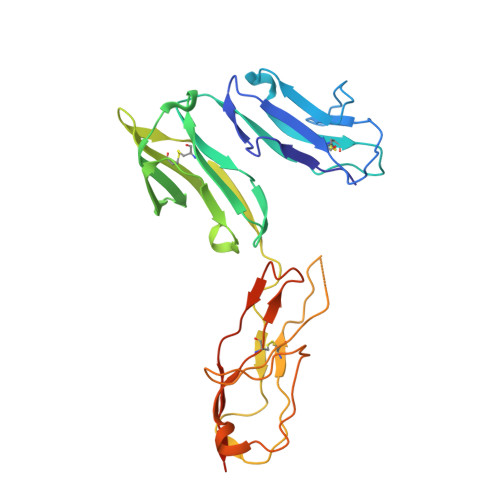
Structure of Fc gamma RI in complex with Fc reveals the importance of glycan recognition for high-affinity IgG binding.
Lu, J., Chu, J., Zou, Z., Hamacher, N.B., Rixon, M.W., Sun, P.D.(2015) Proc Natl Acad Sci U S A 112: 833-838
- PubMed: 25561553 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1418812112
- Primary Citation Related Structures:
4X4M - PubMed Abstract:
Fc gamma receptor I (FcγRI) contributes to protective immunity against bacterial infections, but exacerbates certain autoimmune diseases. The sole high-affinity IgG receptor, FcγRI plays a significant role in immunotherapy. To elucidate the molecular mechanism of its high-affinity IgG binding, we determined the crystal structure of the extracellular domains of human FcγRI in complex with the Fc domain of human IgG1. FcγRI binds to the Fc in a similar mode as the low-affinity FcγRII and FcγRIII receptors. In addition to many conserved contacts, FcγRI forms additional hydrogen bonds and salt bridges with the lower hinge region of Fc. Unique to the high-affinity receptor-Fc complex, however, is the conformation of the receptor D2 domain FG loop, which enables a charged KHR motif to interact with proximal carbohydrate units of the Fc glycans. Both the length and the charge of the FcγRI FG loop are well conserved among mammalian species. Ala and Glu mutations of the FG loop KHR residues showed significant contributions of His-174 and Arg-175 to antibody binding, and the loss of the FG loop-glycan interaction resulted in an ∼ 20- to 30-fold decrease in FcγRI affinity to all three subclasses of IgGs. Furthermore, deglycosylation of IgG1 resulted in a 40-fold loss in FcγRI binding, demonstrating involvement of the receptor FG loop in glycan recognition. These results highlight a unique glycan recognition in FcγRI function and open potential therapeutic avenues based on antibody glycan engineering or small molecular glycan mimics to target FcγRI for certain autoimmune diseases.
- Structural Immunology Section, Laboratory of Immunogenetics, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Rockville, MD 20852 and.
Organizational Affiliation: