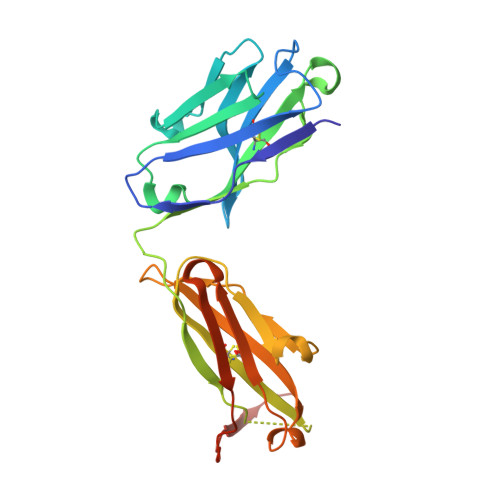
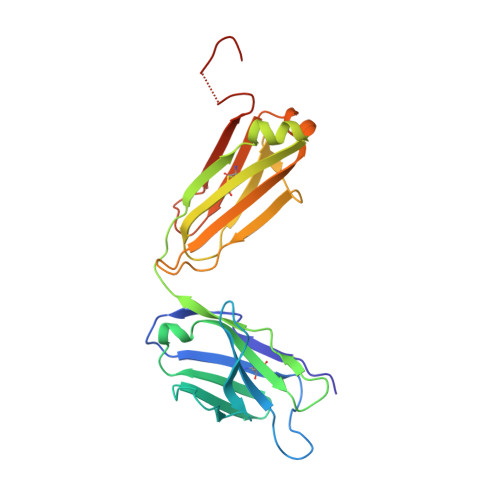
Structural and biophysical characterization of an epitope-specific engineered Fab fragment and complexation with membrane proteins: implications for co-crystallization.
Johnson, J.L., Entzminger, K.C., Hyun, J., Kalyoncu, S., Heaner, D.P., Morales, I.A., Sheppard, A., Gumbart, J.C., Maynard, J.A., Lieberman, R.L.(2015) Acta Crystallogr D Biol Crystallogr 71: 896-906
- PubMed: 25849400 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1399004715001856
- Primary Citation Related Structures:
4X0K - PubMed Abstract:
Crystallization chaperones are attracting increasing interest as a route to crystal growth and structure elucidation of difficult targets such as membrane proteins. While strategies to date have typically employed protein-specific chaperones, a peptide-specific chaperone to crystallize multiple cognate peptide epitope-containing client proteins is envisioned. This would eliminate the target-specific chaperone-production step and streamline the co-crystallization process. Previously, protein engineering and directed evolution were used to generate a single-chain variable (scFv) antibody fragment with affinity for the peptide sequence EYMPME (scFv/EE). This report details the conversion of scFv/EE to an anti-EE Fab format (Fab/EE) followed by its biophysical characterization. The addition of constant chains increased the overall stability and had a negligible impact on the antigen affinity. The 2.0 Å resolution crystal structure of Fab/EE reveals contacts with larger surface areas than those of scFv/EE. Surface plasmon resonance, an enzyme-linked immunosorbent assay, and size-exclusion chromatography were used to assess Fab/EE binding to EE-tagged soluble and membrane test proteins: namely, the β-barrel outer membrane protein intimin and α-helical A2a G protein-coupled receptor (A2aR). Molecular-dynamics simulation of the intimin constructs with and without Fab/EE provides insight into the energetic complexities of the co-crystallization approach.
- School of Chemistry and Biochemistry, Georgia Institute of Technology, 901 Atlantic Drive NW, Atlanta, GA 30332, USA.
Organizational Affiliation: