Redesigning a Monospecific Anti-FGFR3 Antibody to Add Selectivity for FGFR2 and Expand Antitumor Activity.
Yin, Y., Djakovic, S., Marsters, S., Tien, J., Peng, J., Tremayne, J., Lee, G., Neve, R.M., Wu, Y., Merchant, M., Ashkenazi, A., Carter, P.J.(2015) Mol Cancer Ther 14: 2270-2278
- PubMed: 26269606 Search on PubMed
- DOI: https://doi.org/10.1158/1535-7163.MCT-14-1050
- Primary Citation Related Structures:
4WV1 - PubMed Abstract:
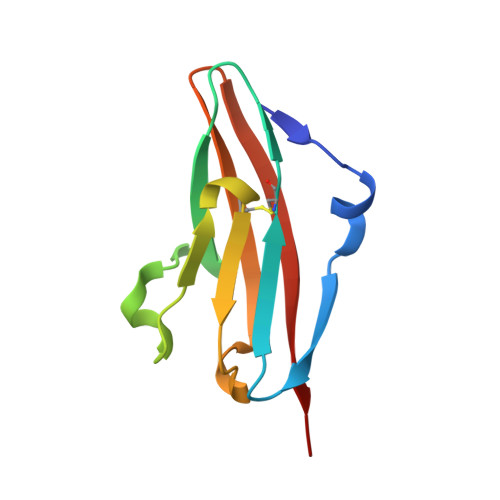
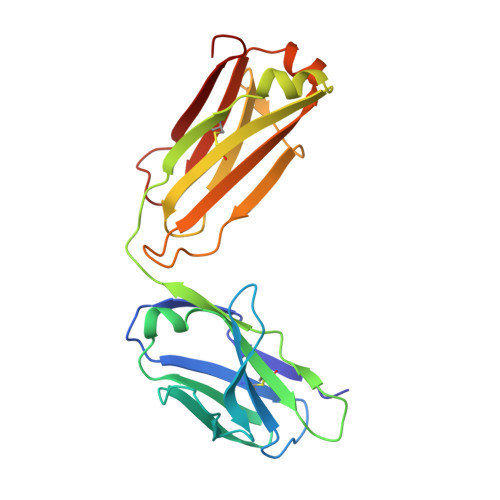
FGF receptors (FGFR) are attractive candidate targets for cancer therapy because they are dysregulated in several human malignancies. FGFR2 and FGFR3 can be inhibited potentially without disrupting adult tissue homeostasis. In contrast, blocking the closely related FGFR1 and FGFR4, which regulate specific metabolic functions, carries a greater safety risk. An anti-FGFR3 antibody was redesigned here to create function-blocking antibodies that bind with dual specificity to FGFR3 and FGFR2 but spare FGFR1 and FGFR4. R3Mab, a previously developed monospecific anti-FGFR3 antibody, was modified via structure-guided phage display and acquired additional binding to FGFR2. The initial variant was trispecific, binding tightly to FGFR3 and FGFR2 and moderately to FGFR4, while sparing FGFR1. The X-ray crystallographic structure indicated that the antibody variant was bound to a similar epitope on FGFR2 as R3Mab on FGFR3. The antibody was further engineered to decrease FGFR4-binding affinity while retaining affinity for FGFR3 and FGFR2. The resulting dual-specific antibodies blocked FGF binding to FGFR3 and FGFR2 and inhibited downstream signaling. Moreover, they displayed efficacy in mice against human tumor xenografts overexpressing FGFR3 or FGFR2. Thus, a monospecific antibody can be exquisitely tailored to confer or remove binding to closely related targets to expand and refine therapeutic potential.
- Department of Antibody Engineering, Genentech, Inc., South San Francisco, California.
Organizational Affiliation: