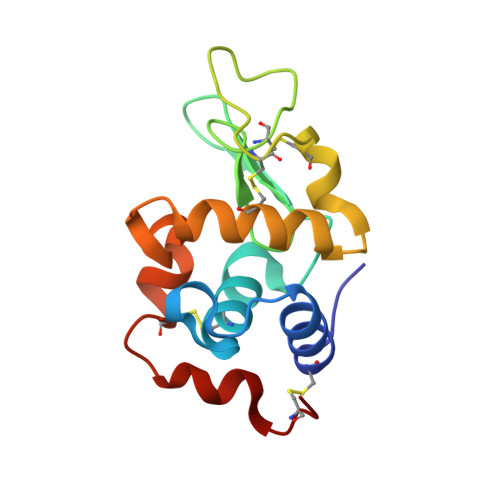
High-pressure protein crystallography of hen egg-white lysozyme
Yamada, H., Nagae, T., Watanabe, N.(2015) Acta Crystallogr D Biol Crystallogr 71: 742-753
- PubMed: 25849385 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1399004715000292
- Primary Citation Related Structures:
4WLD, 4WLT, 4WLX, 4WLY, 4WM1, 4WM2, 4WM3, 4WM4, 4WM5, 4WM6, 4XEN - PubMed Abstract:
Crystal structures of hen egg-white lysozyme (HEWL) determined under pressures ranging from ambient pressure to 950 MPa are presented. From 0.1 to 710 MPa, the molecular and internal cavity volumes are monotonically compressed. However, from 710 to 890 MPa the internal cavity volume remains almost constant. Moreover, as the pressure increases to 950 MPa, the tetragonal crystal of HEWL undergoes a phase transition from P43212 to P43. Under high pressure, the crystal structure of the enzyme undergoes several local and global changes accompanied by changes in hydration structure. For example, water molecules penetrate into an internal cavity neighbouring the active site and induce an alternate conformation of one of the catalytic residues, Glu35. These phenomena have not been detected by conventional X-ray crystal structure analysis and might play an important role in the catalytic activity of HEWL.
- Department of Biotechnology, Graduate School of Engineering, Nagoya University, Chikusa, Nagoya, Aichi 464-8603, Japan.
Organizational Affiliation: