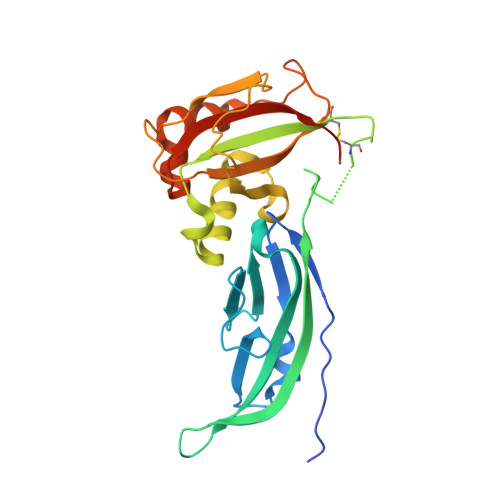
The Structure of the T190M Mutant of Murine alpha-Dystroglycan at High Resolution: Insight into the Molecular Basis of a Primary Dystroglycanopathy.
Bozzi, M., Cassetta, A., Covaceuszach, S., Bigotti, M.G., Bannister, S., Hubner, W., Sciandra, F., Lamba, D., Brancaccio, A.(2015) PLoS One 10: e0124277-e0124277
- PubMed: 25932631 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0124277
- Primary Citation Related Structures:
4WIQ - PubMed Abstract:
The severe dystroglycanopathy known as a form of limb-girdle muscular dystrophy (LGMD2P) is an autosomal recessive disease caused by the point mutation T192M in α-dystroglycan. Functional expression analysis in vitro and in vivo indicated that the mutation was responsible for a decrease in posttranslational glycosylation of dystroglycan, eventually interfering with its extracellular-matrix receptor function and laminin binding in skeletal muscle and brain. The X-ray crystal structure of the missense variant T190M of the murine N-terminal domain of α-dystroglycan (50-313) has been determined, and showed an overall topology (Ig-like domain followed by a basket-shaped domain reminiscent of the small subunit ribosomal protein S6) very similar to that of the wild-type structure. The crystallographic analysis revealed a change of the conformation assumed by the highly flexible loop encompassing residues 159-180. Moreover, a solvent shell reorganization around Met190 affects the interaction between the B1-B5 anti-parallel strands forming part of the floor of the basket-shaped domain, with likely repercussions on the folding stability of the protein domain(s) and on the overall molecular flexibility. Chemical denaturation and limited proteolysis experiments point to a decreased stability of the T190M variant with respect to its wild-type counterpart. This mutation may render the entire L-shaped protein architecture less flexible. The overall reduced flexibility and stability may affect the functional properties of α-dystroglycan via negatively influencing its binding behavior to factors needed for dystroglycan maturation, and may lay the molecular basis of the T190M-driven primary dystroglycanopathy.
- Istituto di Biochimica e Biochimica Clinica, Università Cattolica del Sacro Cuore, Roma 00168, Italy.
Organizational Affiliation: