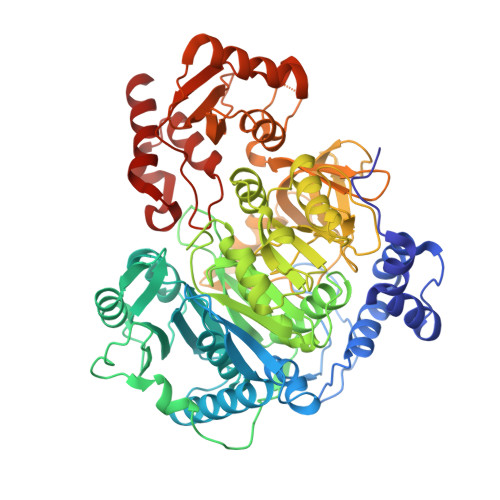
The structure of S. lividans acetoacetyl-CoA synthetase shows a novel interaction between the C-terminal extension and the N-terminal domain.
Mitchell, C.A., Tucker, A.C., Escalante-Semerena, J.C., Gulick, A.M.(2015) Proteins 83: 575-581
- PubMed: 25488501 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.24738
- Primary Citation Related Structures:
4WD1 - PubMed Abstract:
The adenosine monoposphate-forming acyl-CoA synthetase enzymes catalyze a two-step reaction that involves the initial formation of an acyl adenylate that reacts in a second partial reaction to form a thioester between the acyl substrate and CoA. These enzymes utilize a Domain Alternation catalytic mechanism, whereby a ∼ 110 residue C-terminal domain rotates by 140° to form distinct catalytic conformations for the two partial reactions. The structure of an acetoacetyl-CoA synthetase (AacS) is presented that illustrates a novel aspect of this C-terminal domain. Specifically, several acetyl- and acetoacetyl-CoA synthetases contain a 30-residue extension on the C-terminus compared to other members of this family. Whereas residues from this extension are disordered in prior structures, the AacS structure shows that residues from this extension may interact with key catalytic residues from the N-terminal domain.
- Hauptman-Woodward Institute, Buffalo, New York, 14203; Department of Structural Biology, University at Buffalo, Buffalo, New York, 14203.
Organizational Affiliation: