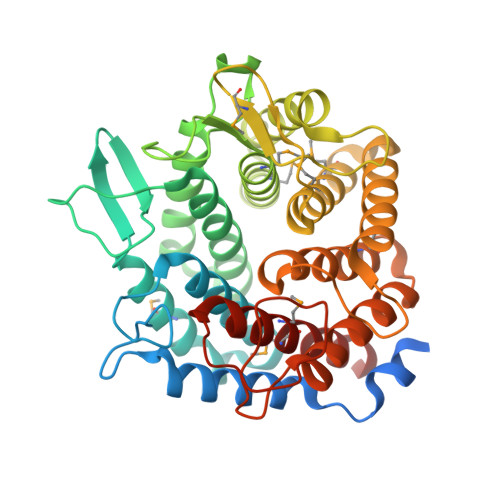
Structure of the Gh76 Alpha-Mannanase Homolog, Bt2949, from the Gut Symbiont Bacteroides Thetaiotaomicron
Thompson, A.J., Cuskin, F., Spears, R.J., Dabin, J., Turkenburg, J.P., Gilbert, H.J., Davies, G.J.(2015) Acta Crystallogr D Biol Crystallogr 71: 408
- PubMed: 25664752 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1399004714026443
- Primary Citation Related Structures:
4V1R, 4V1S - PubMed Abstract:
The large bowel microbiota, a complex ecosystem resident within the gastrointestinal tract of all human beings and large mammals, functions as an essential, nonsomatic metabolic organ, hydrolysing complex dietary polysaccharides and modulating the host immune system to adequately tolerate ingested antigens. A significant member of this community, Bacteroides thetaiotaomicron, has evolved a complex system for sensing and processing a wide variety of natural glycoproducts in such a way as to provide maximum benefit to itself, the wider microbial community and the host. The immense ability of B. thetaiotaomicron as a `glycan specialist' resides in its enormous array of carbohydrate-active enzymes, many of which are arranged into polysaccharide-utilization loci (PULs) that are able to degrade sugar polymers that are often inaccessible to other gut residents, notably α-mannan. The B. thetaiotaomicron genome encodes ten putative α-mannanases spread across various PULs; however, little is known about the activity of these enzymes or the wider implications of α-mannan metabolism for the health of both the microbiota and the host. In this study, SAD phasing of a selenomethionine derivative has been used to investigate the structure of one such B. thetaiotaomicron enzyme, BT2949, which belongs to the GH76 family of α-mannanases. BT2949 presents a classical (α/α)6-barrel structure comprising a large extended surface cleft common to other GH76 family members. Analysis of the structure in conjunction with sequence alignments reveals the likely location of the catalytic active site of this noncanonical GH76.
- Department of Chemistry, University of York, Heslington, York YO10 5DD, England.
Organizational Affiliation: