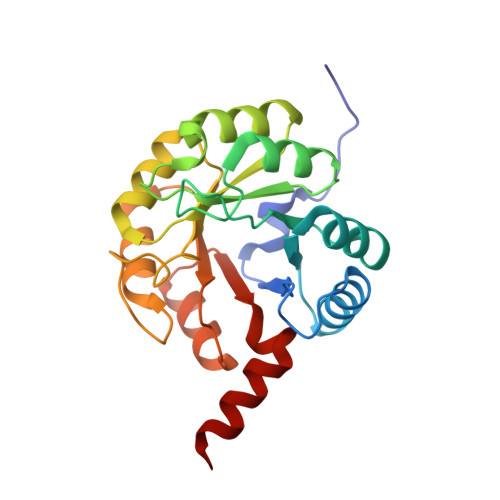
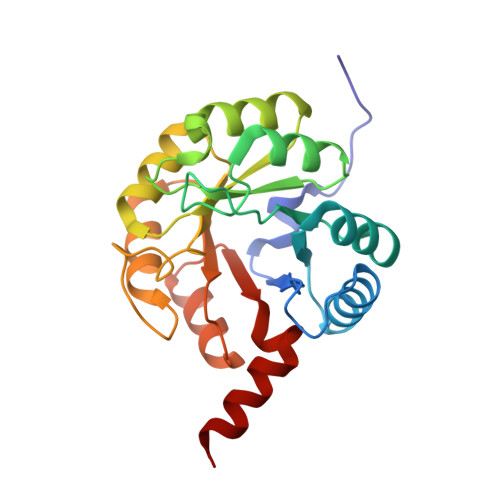
Structural and Functional Characterization of the Clostridium Perfringens N-Acetylmannosamine-6-Phosphate 2-Epimerase Essential for the Sialic Acid Salvage Pathway.
Pelissier, M., Sebban-Kreuzer, C., Guerlesquin, F., Brannigan, J.A., Bourne, Y., Vincent, F.(2014) J Biological Chem 289: 35215
- PubMed: 25320079 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.604272
- Primary Citation Related Structures:
4UTT, 4UTU, 4UTW - PubMed Abstract:
Pathogenic bacteria are endowed with an arsenal of specialized enzymes to convert nutrient compounds from their cell hosts. The essential N-acetylmannosamine-6-phosphate 2-epimerase (NanE) belongs to a convergent glycolytic pathway for utilization of the three amino sugars, GlcNAc, ManNAc, and sialic acid. The crystal structure of ligand-free NanE from Clostridium perfringens reveals a modified triose-phosphate isomerase (β/α)8 barrel in which a stable dimer is formed by exchanging the C-terminal helix. By retaining catalytic activity in the crystalline state, the structure of the enzyme bound to the GlcNAc-6P product identifies the topology of the active site pocket and points to invariant residues Lys(66) as a putative single catalyst, supported by the structure of the catalytically inactive K66A mutant in complex with substrate ManNAc-6P. (1)H NMR-based time course assays of native NanE and mutated variants demonstrate the essential role of Lys(66) for the epimerization reaction with participation of neighboring Arg(43), Asp(126), and Glu(180) residues. These findings unveil a one-base catalytic mechanism of C2 deprotonation/reprotonation via an enolate intermediate and provide the structural basis for the development of new antimicrobial agents against this family of bacterial 2-epimerases.
- From the Aix-Marseille University, AFMB UMR7257, 163 avenue de Luminy 13288 Marseille, France, the CNRS, AFMB UMR7257, 163 avenue de Luminy, 13288 Marseille, France.
Organizational Affiliation: