Insights Into the Bioactivity of Mensacarcin and Epoxide Formation by Msno8.
Maier, S., Pflueger, T., Loesgen, S., Asmus, K., Broetz, E., Paululat, T., Zeeck, A., Andrade, S., Bechthold, A.(2014) Chembiochem 15: 749
- PubMed: 24554499 Search on PubMed
- DOI: https://doi.org/10.1002/cbic.201300704
- Primary Citation Related Structures:
4US5 - PubMed Abstract:
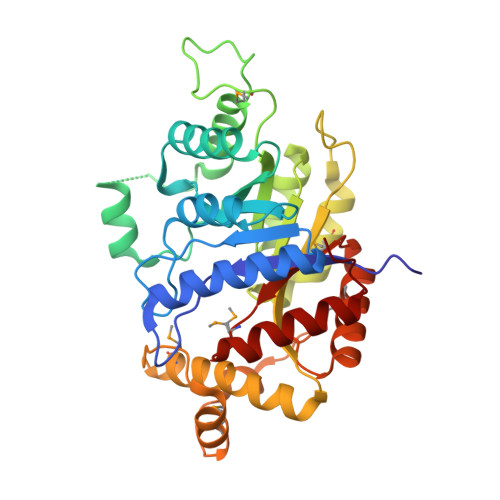
Mensacarcin, a potential antitumour drug, is produced by Streptomyces bottropensis. The structure consists of a three-membered ring system with many oxygen atoms. Of vital importance in this context is an epoxy moiety in the side chain of mensacarcin. Our studies with different mensacarcin derivatives have demonstrated that this epoxy group is primarily responsible for the cytotoxic effect of mensacarcin. In order to obtain further information about this epoxy moiety, inactivation experiments in the gene cluster were carried out to identify the epoxy-forming enzyme. Therefore the cosmid cos2, which covers almost the complete type II polyketide synthase (PKS) gene cluster, was heterologously expressed in Streptomyces albus. This led to production of didesmethylmensacarcin, due to the fact that methyltransferase genes are missing in the cosmid. Further gene inactivation experiments on this cosmid showed that MsnO8, a luciferase-like monooxygenase, introduces the epoxy group at the end of the biosynthesis of mensacarcin. In addition, the protein MsnO8 was purified, and its crystal structure was determined to a resolution of 1.80 Å.
- Institut für Pharmazeutische Biologie und Biotechnologie, Albert-Ludwigs Universität, Stefan-Meier-Strasse 19, 79104 Freiburg (Germany).
Organizational Affiliation: