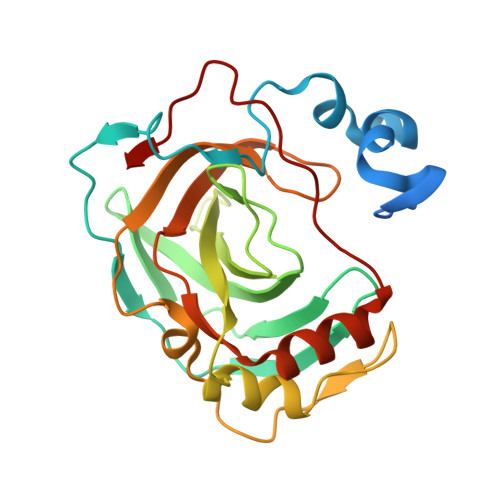
The Structure of a Tetrameric [Alpha]-Carbonic Anhydrase from Thermovibrio Ammonificans Reveals a Core Formed Around Intermolecular Disulfides that Contribute to its Thermostability
James, P., Isupov, M.N., Sayer, C., Saneei, V., Berg, S., Lioliou, M., Kotlar, H., Littlechild, J.(2014) Acta Crystallogr D Biol Crystallogr 70: 2607
- PubMed: 25286845 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004714016526
- Primary Citation Related Structures:
4C3T, 4COQ, 4UOV - PubMed Abstract:
Carbonic anhydrase enzymes catalyse the reversible hydration of carbon dioxide to bicarbonate. A thermophilic Thermovibrio ammonificans α-carbonic anhydrase (TaCA) has been expressed in Escherichia coli and structurally and biochemically characterized. The crystal structure of TaCA has been determined in its native form and in two complexes with bound inhibitors. The tetrameric enzyme is stabilized by a unique core in the centre of the molecule formed by two intersubunit disulfides and a single lysine residue from each monomer that is involved in intersubunit ionic interactions. The structure of this core protects the intersubunit disulfides from reduction, whereas the conserved intrasubunit disulfides are not formed in the reducing environment of the E. coli host cytosol. When oxidized to mimic the environment of the periplasmic space, TaCA has increased thermostability, retaining 90% activity after incubation at 70°C for 1 h, making it a good candidate for industrial carbon-dioxide capture. The reduction of all TaCA cysteines resulted in dissociation of the tetrameric molecule into monomers with lower activity and reduced thermostability. Unlike other characterized α-carbonic anhydrases, TaCA does not display esterase activity towards p-nitrophenyl acetate, which appears to result from the increased rigidity of its protein scaffold.
- Henry Wellcome Building for Biocatalysis, Biosciences, College of Life and Environmental Sciences, University of Exeter, Stocker Road, Exeter EX4 4QD, England.
Organizational Affiliation: