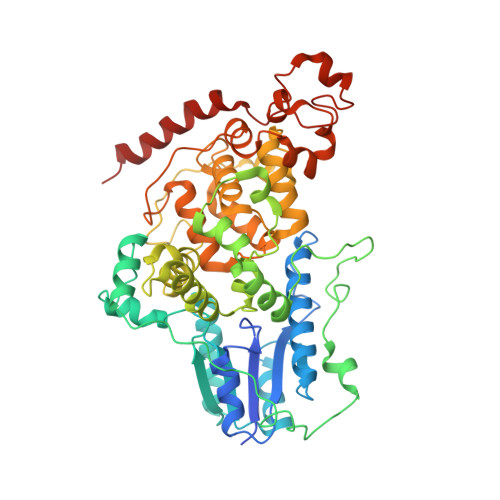
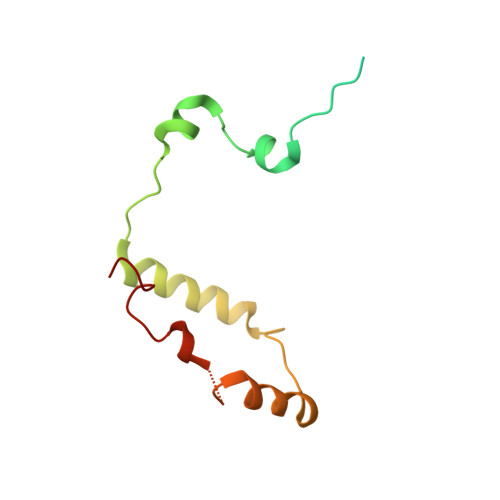
Molecular assembly of the period-cryptochrome circadian transcriptional repressor complex.
Nangle, S.N., Rosensweig, C., Koike, N., Tei, H., Takahashi, J.S., Green, C.B., Zheng, N.(2014) Elife 3: e03674-e03674
- PubMed: 25127877 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.03674
- Primary Citation Related Structures:
4U8H - PubMed Abstract:
The mammalian circadian clock is driven by a transcriptional-translational feedback loop, which produces robust 24-hr rhythms. Proper oscillation of the clock depends on the complex formation and periodic turnover of the Period and Cryptochrome proteins, which together inhibit their own transcriptional activator complex, CLOCK-BMAL1. We determined the crystal structure of the CRY-binding domain (CBD) of PER2 in complex with CRY2 at 2.8 Å resolution. PER2-CBD adopts a highly extended conformation, embracing CRY2 with a sinuous binding mode. Its N-terminal end tucks into CRY adjacent to a large pocket critical for CLOCK-BMAL1 binding, while its C-terminal half flanks the CRY2 C-terminal helix and sterically hinders the recognition of CRY2 by the FBXL3 ubiquitin ligase. Unexpectedly, a strictly conserved intermolecular zinc finger, whose integrity is important for clock rhythmicity, further stabilizes the complex. Our structure-guided analyses show that these interspersed CRY-interacting regions represent multiple functional modules of PERs at the CRY-binding interface.
- Department of Pharmacology, University of Washington, Seattle, United States.
Organizational Affiliation: