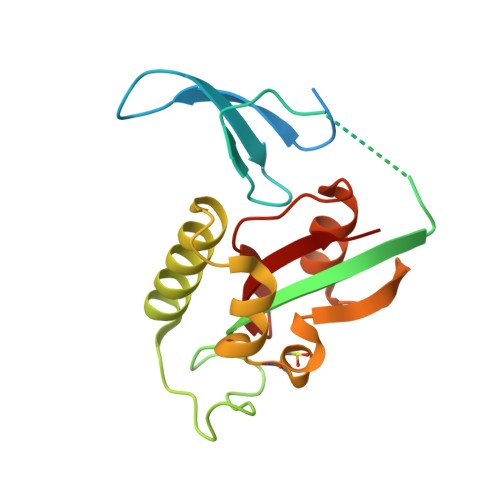
Pin1 cysteine-113 oxidation inhibits its catalytic activity and cellular function in Alzheimer's disease.
Chen, C.H., Li, W., Sultana, R., You, M.H., Kondo, A., Shahpasand, K., Kim, B.M., Luo, M.L., Nechama, M., Lin, Y.M., Yao, Y., Lee, T.H., Zhou, X.Z., Swomley, A.M., Allan Butterfield, D., Zhang, Y., Lu, K.P.(2015) Neurobiol Dis 76: 13-23
- PubMed: 25576397 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.nbd.2014.12.027
- Primary Citation Related Structures:
4U84, 4U85, 4U86 - PubMed Abstract:
The unique proline isomerase Pin1 is pivotal for protecting against age-dependent neurodegeneration in Alzheimer's disease (AD), with its inhibition providing a molecular link between tangle and plaque pathologies. Pin1 is oxidatively modified in human AD brains, but little is known about its regulatory mechanisms and pathological significance of such Pin1 modification. In this paper, our determination of crystal structures of oxidized Pin1 reveals a series of Pin1 oxidative modifications on Cys113 in a sequential fashion. Cys113 oxidization is further confirmed by generating antibodies specifically recognizing oxidized Cys113 of Pin1. Furthermore, Pin1 oxidation on Cys113 inactivates its catalytic activity in vitro, and Ala point substitution of Cys113 inactivates the ability of Pin1 to isomerize tau as well as to promote protein turnover of tau and APP. Moreover, redox regulation affects Pin1 subcellular localization and Pin1-mediated neuronal survival in response to hypoxia treatment. Importantly, Cys113-oxidized Pin1 is significantly increased in human AD brain comparing to age-matched controls. These results not only identify a novel Pin1 oxidation site to be the critical catalytic residue Cys113, but also provide a novel oxidative regulation mechanism for inhibiting Pin1 activity in AD. These results suggest that preventing Pin1 oxidization might help to reduce the risk of AD.
- Department of Medicine, Beth Israel Deaconess Medical Center, Harvard Medical School, Boston, MA 02215, USA.
Organizational Affiliation: