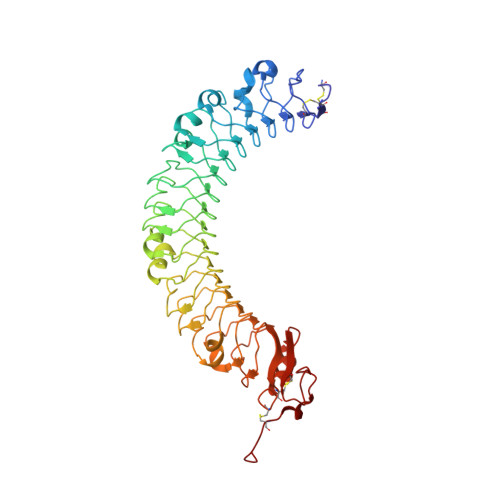
LRIG1 Extracellular Domain: Structure and Function Analysis.
Xu, Y., Soo, P., Walker, F., Zhang, H.H., Redpath, N., Tan, C.W., Nicola, N.A., Adams, T.E., Garrett, T.P., Zhang, J.G., Burgess, A.W.(2015) J Mol Biology 427: 1934-1948
- PubMed: 25765764 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2015.03.001
- Primary Citation Related Structures:
4U7L, 4U7M - PubMed Abstract:
We have expressed and purified three soluble fragments of the human LRIG1-ECD (extracellular domain): the LRIG1-LRR (leucine-rich repeat) domain, the LRIG1-3Ig (immunoglobulin-like) domain, and the LRIG1-LRR-1Ig fragment using baculovirus vectors in insect cells. The two LRIG1 domains crystallised so that we have been able to determine the three-dimensional structures at 2.3Å resolution. We developed a three-dimensional structure for the LRIG1-ECD using homology modelling based on the LINGO-1 structure. The LRIG1-LRR domain and the LRIG1-LRR-1Ig fragment are monomers in solution, whereas the LRIG1-3Ig domain appears to be dimeric. We could not detect any binding of the LRIG1 domains or the LRIG1-LRR-1Ig fragment to the EGF receptor (EGFR), either in solution using biosensor analysis or when the EGFR was expressed on the cell surface. The FLAG-tagged LRIG1-LRR-1Ig fragment binds weakly to colon cancer cells regardless of the presence of EGFRs. Similarly, neither the soluble LRIG1-LRR nor the LRIG1-3Ig domains nor the full-length LRIG1 co-expressed in HEK293 cells inhibited ligand-stimulated activation of cell-surface EGFR.
- Structural Biology Division, The Walter and Eliza Hall Institute of Medical Research, Parkville, Victoria 3052, Australia; Cancer and Haematology Division, The Walter and Eliza Hall Institute of Medical Research, Parkville, Victoria 3052, Australia.
Organizational Affiliation: